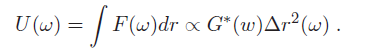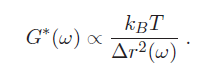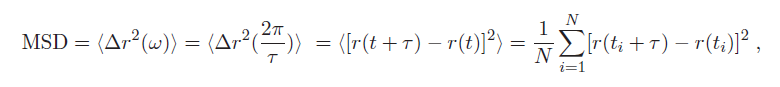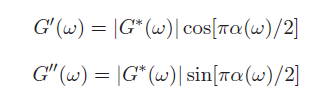Cellular microrheology
Introduction and Background
Microrheology
Many cellular functions such as migration, differentiation, and proliferation are regulated by the mechanical properties of cells, specifically, their elasticity and viscosity. Rheology is the science of measuring materials' mechanical properties. Microrheology is a subgroup of techniques that are capable of measuring mechanical properties from microscopic material volumes. Clearly, given the typical size of biological cells, microrheology is the technique needed to measure their elasticity and viscosity. The elastic and viscous properties of cells can be characterized by a complex-valued shear modulus (with units of Pa) G*(ω) = G'(ω) + iG"(ω). The real part G'(ω), referred to as the storage modulus, is a measure of cell elasticity, while the imaginary part G"(ω), the loss modulus, is a measure of their viscosity. A generalized Hookean relationship can be written as:
where Δr is a generalized displacement, and F(ω) is a force linearly proportional to it via the shear modulus. Therefore, we can measure the shear modulus if we can measure the deformation of the cell under a known force. (Note that all these quantities are frequency-dependent). Particle-tracking microrheometry is based on measuring the displacement of a particle with radius $ a $ embedded in a cell driven by thermal forces. One complication is that this relationship is frequency dependent - this is because in complex fluids, such as the cellular cytoskeleton, there are different energy dissipation mechanisms over different time scales. To approach the derivation of the relevant formulas, it is more convenient to think in terms of energy, rather than force. The relationship between stored energy and displacement has a familiar form, similar to a spring-mass system (recall $ KE \ \alpha \ k(\omega)^2 $ ):
What is the driving thermal energy U(ω)? Recall also that thermal energy is "white," i.e. it contains equal power at all frequencies and is equal to 0.5*k$ _B $T for each degree of freedom in a second-order system, where k$ _B $ is Boltzmann's constant and T is the absolute temperature. From this relationship (since we're observing motion in two dimensions), we have
Our argument is obviously very approximate. A complete (and much more difficult) derivation results in the following equation (see Mason [1] for details):
$ \left\vert G^*(\omega) \right\vert = {k_B T \over \pi a \left \langle \Delta r^2 (1/\omega) \right \rangle \Gamma[1 + \alpha(\omega)]} $
Some key additional details to help you make sense of this equation:
1. As you can see, the dependence on displacement is more accurately expressed as the mean squared displacement (MSD) <Δr$ ^2 $(1/ω)>:
where < > denotes a time-average of the particle's displacement trajectory r(t), at discrete times t = t$ _1 $,.... t$ _n $ (as sampled by a digital system like the PC and camera). Additionally, τ is a characteristic lag/delay time for the measurement. corresponding to the frequency ω.
2. $ \alpha(\omega)=\frac{\partial \ln \left \langle \Delta r^2(\tau) \right \rangle }{\partial \ln \tau} $
3. The radius of the particle $ a $ plays a role in the formula.
4. Γ(.) is the Gamma function (the generalized form of the factorial function, which can be looked up in a mathematical table). Mason suggests that for our range of α,
$ \Gamma[1 + \alpha] \approx = 0.457(1 + \alpha)^2 - 1.36(1 + \alpha) + 1.90. $
This equation may look complicated but there is a simple approximation to calculate the elastic and viscous moduli:
A detailed discussion of particle tracking microrheology can be found in the papers by Mason and Lau[2]. An application of the work of Mason et al. to 3T3 cells in particular can be found in the work of Tseng et al. [3]
Optical microscopy lab
Code examples and simulations
- Converting Gaussian fit to Rayleigh resolution
- MATLAB: Estimating resolution from a PSF slide image
- Matlab: Scalebars
- Calculating MSD and Diffusion Coefficients
Background reading
- Geometrical optics and ray tracing
- Physical optics and resolution
- Optical aberrations
- Aperture and field stops
- Optical detectors, noise, and the limit of detection
- Manta G032 camera measurements
- Understanding log plots
References
- ↑ T. G. Mason, "Estimating the viscoelastic moduli of complex fluids using the generalized Stokes-Einstein equation" Rheol. Acta, 39, pp. 371-378 (2000).
- ↑ A. W. C. Lau et al., "Microrheology, Stress Fluctuations, and Active Behavior of Living Cells," Phys. Rev. Lett., 91(19), p. 198101 (2003)
- ↑ Yiider Tseng, Thomas P Kole, and Denis Wirtz, "Micromechanical mapping of live cells by multiple-particle-tracking microrheology." Biophys J. 2002 December; 83(6): 3162–3176.