Difference between revisions of "20.109(S17):Complete cell viability assay (Day9)"
(→Part 1: Evaluate cell viability) |
(→Part 1: Evaluate cell viability) |
||
| (5 intermediate revisions by one user not shown) | |||
| Line 24: | Line 24: | ||
#*Be very careful that you add your samples to the appropriate wells according the plate map below. | #*Be very careful that you add your samples to the appropriate wells according the plate map below. | ||
#*Instructors will add media only control to the 96-well plate to obtain a value for background luminescence. | #*Instructors will add media only control to the 96-well plate to obtain a value for background luminescence. | ||
| − | #When all teams have transferred their samples, the teaching faculty will | + | #When all teams have transferred their samples, the teaching faculty will measure the luminescence using a Molecular Devices SpectraMax M3 Microplate Reader. |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
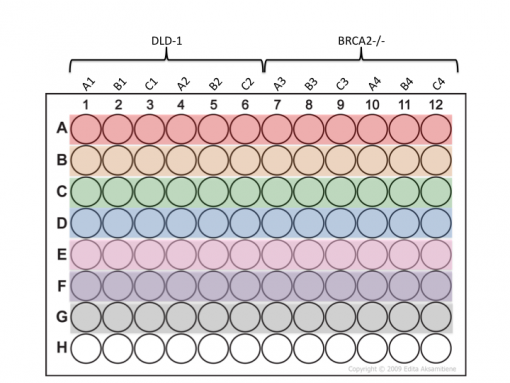
[[File:Sp18 20.109 CTG plate map.png|thumb|510px|center|]] | [[File:Sp18 20.109 CTG plate map.png|thumb|510px|center|]] | ||
| Line 35: | Line 30: | ||
===Part 2: Complete data analysis=== | ===Part 2: Complete data analysis=== | ||
Pool the data by drug and use the statistical tools you learned in Mod1 to analyze the relevant class data for your M2 Research Article. | Pool the data by drug and use the statistical tools you learned in Mod1 to analyze the relevant class data for your M2 Research Article. | ||
| + | |||
| + | #Review your data posted on the [[20.109(S18):Class data| Class data page]]. | ||
| + | #The data are represented as raw luminescent vales (arbitrary units). | ||
| + | #Subtract the average of the three ‘no cells control’ from all of the luminescent vales. This is the background luminescence. | ||
| + | #Normalize your luminescence values by dividing all wells by the ‘no DNA damage controls’ value for the appropriate cell line. This gives you viability as the “% of control.” | ||
| + | # You can average class data to include error bars and information concerning statistical significance. | ||
==Reagents== | ==Reagents== | ||
*CellTiter Glo cell viability assay kit (Promega) | *CellTiter Glo cell viability assay kit (Promega) | ||
| + | *SpectraMax M3 Microplate Reader (Molecular Devices) | ||
==Navigation links== | ==Navigation links== | ||
Next day: [[20.109(S18):Growth of phage-based active (cathode) material (Day1) | Grow phage-based active (cathode) material]]<br> | Next day: [[20.109(S18):Growth of phage-based active (cathode) material (Day1) | Grow phage-based active (cathode) material]]<br> | ||
Previous day: [[20.109(S18):Induce DNA damage and apply drug treatments for cell viability and identification of regulatory motifs in RNA-seq data (Day8)| Induce DNA damage and apply drug treatments for cell viability and identification of regulatory motifs in RNA-seq data]] | Previous day: [[20.109(S18):Induce DNA damage and apply drug treatments for cell viability and identification of regulatory motifs in RNA-seq data (Day8)| Induce DNA damage and apply drug treatments for cell viability and identification of regulatory motifs in RNA-seq data]] | ||
Latest revision as of 15:29, 13 April 2018
Contents
Introduction
The CellTiter-Glo Luminescent Cell Viability Assay is a method for quantifying the number of viable cells based on measuring the amount of ATP present. ATP is a proxy for the presence of metabolically active (alive) cells. In this assay, the cells are lysed and ATP is released from the active cells. In a reaction catalyzed by a propriety luciferase enzyme, luciferin, ATP, and oxygen result in oxyluciferin, AMP, PPi, carbon dioxide, and light. The light product is then measured using a luminometer.
Protocols
Part 1: Evaluate cell viability
Today you will complete the second experiment for Mod 2 - the cell viability assay. In the previous class you used etoposide to induce DNA damage and added a NHEJ inhibitor. By comparing the results of these treatments between the DLD-1 and BRCA2- cell lines you will more deeply examine the DNA repair pathways from lecture.
- Retrieve your plate from the 37 °C incubator.
- Briefly look at the DLD1 and BRCA2-/- cells under the microscope.
- Make a note of their confluency and morphology.
- Aspirate the spent media from each well.
- Be careful not to cross-contaminate between the wells.
- Add 500 μL of fresh media into each well.
- Obtain an aliquot of the CellTiter Glo reagent from the front laboratory bench.
- Thoroughly mix the CellTiter Glo reagent, then add 500 μL into each well of your plate.
- After adding the reagent, pipet up and down ten times to mix.
- Move your plate to the plate shaker at the front laboratory bench and shake for 2 min.
- Remove your plate from the plate shaker and incubate on your benchtop for 10 min.
- Transfer 100 μL from each well into a white 96-well plate.
- Be very careful that you add your samples to the appropriate wells according the plate map below.
- Instructors will add media only control to the 96-well plate to obtain a value for background luminescence.
- When all teams have transferred their samples, the teaching faculty will measure the luminescence using a Molecular Devices SpectraMax M3 Microplate Reader.
Part 2: Complete data analysis
Pool the data by drug and use the statistical tools you learned in Mod1 to analyze the relevant class data for your M2 Research Article.
- Review your data posted on the Class data page.
- The data are represented as raw luminescent vales (arbitrary units).
- Subtract the average of the three ‘no cells control’ from all of the luminescent vales. This is the background luminescence.
- Normalize your luminescence values by dividing all wells by the ‘no DNA damage controls’ value for the appropriate cell line. This gives you viability as the “% of control.”
- You can average class data to include error bars and information concerning statistical significance.
Reagents
- CellTiter Glo cell viability assay kit (Promega)
- SpectraMax M3 Microplate Reader (Molecular Devices)
Next day: Grow phage-based active (cathode) material