20.109(S18):Module 2
Contents
Module 2
Lecturers: Ernest Fraenkel and Noreen Lyell
Instructors: Noreen Lyell, Leslie McClain and Josephine Bagnall
TA: Casper Enghuus
Lab manager: Hsinhwa Lee
Overview
Non-homologous end joining (NHEJ) is a pathway of DNA repair by which double-strand breaks are ligated together. In specific types of cancer, targeting NHEJ is an effective treatment option as a paradox exists in cancer. Whole genome sequencing has revealed that the cells of many tumor types have mutations in genes necessary for DNA repair. These mutations are responsible for cells becoming cancerous, but are also detrimental because, just like normal cells, cancer cells must divide to survive. Thus, a cancer cell will develop an ‘addiction’ to a DNA repair pathway – specifically, a pathway different from the one with the mutation that caused the cell to generate a tumor. Recent cancer therapies seek to exploit this addiction by targeting the intact pathway used by the tumor cells to repair DNA damage due to intrinsic breaks that occur during replication. In addition, the effectiveness of DNA damage induced by chemotherapy treatment may be enhanced by also disrupting the functional repair pathways of tumor cells.
In this module, you will examine the effect of DNA damage on cells that are deficient in homologous recombination, a DNA repair pathway that uses sequence homology to correct double-strand breaks. To this end, you will address two primary research questions:
- What genes are differentially expressed in response to DNA damage and drug treatment?
- Does drug treatment affect cell survival following DNA damage?
An additional goal of Module 2 is for you to synthesize data from different experiments into a single story. More often that not, researchers reach conclusions using results gathered from more than one experimental approach. Multiple layers of support can bolster a conclusion as they eliminate biases that may exist in a particular method of analysis. Furthermore, for complex questions it may be necessary to include data from several approaches to provide a sufficient conclusion.
Lab links: day by day
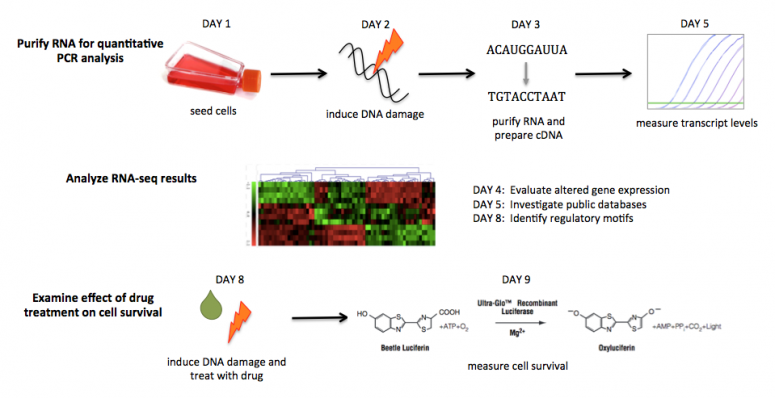
M2D1: Practice tissue culture techniques and prepare cells for RNA purification
M2D2: Induce DNA damage and apply drug treatments for RNA purification
M2D3: Purify RNA and practice RNA-seq data analysis methods
M2D4: Analyze RNA-seq data and prepare for quantitative PCR experiment
M2D5: Investigate RNA-seq data using public databases
M2D6: Journal club I presentations
M2D7: Journal club II presentations
M2D8: Induce DNA damage and apply drug treatments for cell viability and identification of regulatory motifs in RNA-seq data
M2D9: Complete cell viability assay
Assignments
Journal club presentation
Research article
References
- A syngeneic variance library for functional annotation of human variation: application to BRCA2. Cancer Research. 68:5023-5030.
- Synthetic lethality: exploiting the addiction of cancer to DNA repair. Blood Journal. 117:6074-6082.