20.109(S18):Module 1
Contents
Module 1
Lecturer: Angela Koehler
Instructors: Noreen Lyell, Leslie McClain and Josephine Bagnall
TA: Casper Enghuus
Lab manager: Hsinhwa Lee
Overview
Chemical probes, or ligands, are important research tools used to explore cellular processes and therapeutic targets. The use of high-throughput and unbiased strategies to identify small molecules that bind specific biomolecules, such as proteins, can provide insight on the structure or function of targets. Additionally, a small-molecule screen can identify new chemical probes for target proteins of interest.
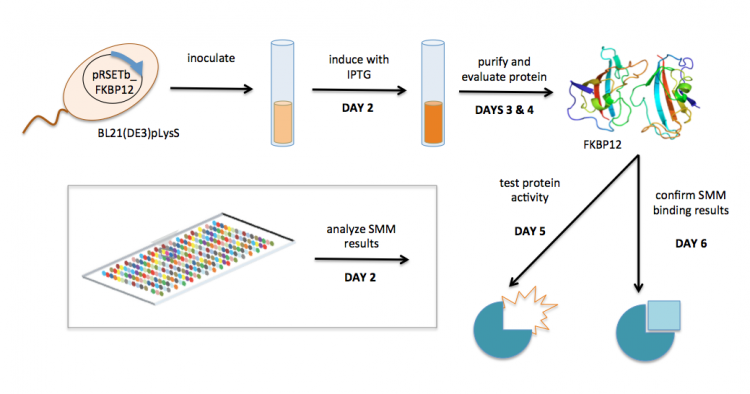
The small-molecule microarray (SMM) is a high-throughput method that enables the detection of protein-ligand binding. Briefly, ligands are 'printed' onto a slide and incubated with purifed protein. Unbound protein is washed from the slide and bound protein is detected using a tag on the protein of interest. Because the location of every ligand on the slide is known, the detection of protein indicates that it is bound to the ligand at that location.
In your experiment, you will use SMM data collected by students from the Sp17 semester to identify ligands that bind to FKBP12, a folding chaperone for proteins that contain proline residues in eukaryotes. To expand on the research completed by the previous 109ers, you will confirm that the identified ligands bind FKBP12 using secondary assays.
This module has been developed thanks to the generous time and thoughtful efforts of several Koehler Laboratory members, in particular Rob Wilson, Dr. Becky Leifer, and Shelby Doyle.
Lab links: day by day
M1D1: In silico cloning and confirmation digest of protein expression vector
M1D2: Complete small microarray analysis and induce protein expression
M1D3: Purify protein for secondary assays
M1D4: Evaluate protein purity and concentration
M1D5: Test protein activity using peptidyl-prolyl cis-trans isomerase assay
M1D6: Confirm ligand binding using differential scanning fluorimetry assay
M1D7: Complete data analysis
Assignments
Data summary
Mini-presentation
References
- A method for the covalent capture and screening of diverse small molecules in a microarray format. Nature Protocols. 1:2344-2352.
- Recent discoveries and applications involving small-molecule microarrays. Chemical Biology. 18:21-28.