Difference between revisions of "20.109(S17):Examine transcript levels in response to DNA damage (Day 4)"
Noreen Lyell (Talk | contribs) (→Introduction) |
Noreen Lyell (Talk | contribs) (→Part 1: Setup quantitative PCR assay) |
||
| (One intermediate revision by one user not shown) | |||
| Line 17: | Line 17: | ||
===Part 1: Setup quantitative PCR assay=== | ===Part 1: Setup quantitative PCR assay=== | ||
| + | |||
| + | [[Image:Sp17 20.109 M2D2 qPCR.png|thumb|750px|center|]] | ||
| + | #*Label 8 microcentrifuge tubes according to the samples you will prepare for RRT-PCR. | ||
| + | #*Obtain the cDNA aliquots from the front lab bench (sample #1, #2, #3). | ||
| + | #*Each reaction should contain 4 μL of cDNA (at 6.25 ng/uL), 2 μL of your diluted primer solution, and water for a total volume of 12.5 μL. For each sample/primer set combination, prepare enough for 2.5 reactions. | ||
| + | #Add your reactions to the plate on the front bench according to the map. You will add 12.5 μL to each well. | ||
| + | #Once everyone has added their reactions, we will add 12.5 μL of the 2X Syber Green master mix to each well. | ||
| + | |||
| + | For your reference, the PCR cycling conditions will be those shown below: | ||
| + | |||
| + | <center> | ||
| + | {| border="1" | ||
| + | ! Stage | ||
| + | ! Cycles | ||
| + | ! Details | ||
| + | |- | ||
| + | | 1 | ||
| + | | 1 | ||
| + | | 95°C for 10 min | ||
| + | |- | ||
| + | | 2 | ||
| + | | 40 | ||
| + | | 95°C for 15 sec <br> 60°C for 30 sec <br> 72°C for 30 sec | ||
| + | |- | ||
| + | | 3 | ||
| + | | 1 | ||
| + | | 4°C hold | ||
| + | |} | ||
| + | </center> | ||
===Part 2: Discussion of Research article details=== | ===Part 2: Discussion of Research article details=== | ||
Latest revision as of 02:33, 3 February 2017
Contents
Introduction
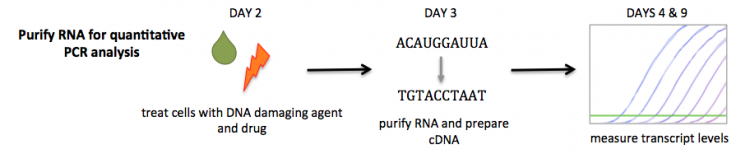
Real-time reverse transcription-polymerase chain reaction (RRT-PCR) allows researchers to monitor the results of PCR as amplification is occurring (this technique is also referred to as quantitative or real-time PCR). During RRT-PCR data are collected throughout the amplification process using a fluorescent dye. The fluorescent dye is highly specific for double-stranded DNA and when bound to DNA molecules the fluorescence intensity increases proportionate to the increase in double-stranded product. In contrast, the data for traditional PCR are simply observed as a band on a gel (remember back to M1D5).
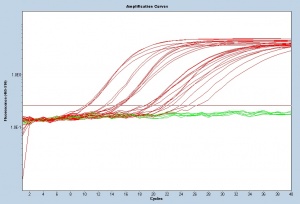
To compare the sensitivity of your primers to the sensitivity of those currently used in the Runstadler lab you will examine the CT values from your RRT-PCR assay. The CT values are displayed as an amplification curve following RRT-PCR (these values are also given numerically). The initial cycles measure very little fluorescence due to low amounts of double-stranded DNA and are used to establish the inherent background fluorescence. As double-stranded product is produced, fluorescence is measured and the curve appears linear. This linear portion of the curve represents the exponential phase of PCR. Throughout the exponential phase, the curve should be smooth. Sharp points may be due to errors in reaction preparation or failures in the machine used to measure fluorescence. As mentioned previously, the first cycle in which the fluorescence measurement is above background is the CT. During the later cycles the curve shows minimal increases in fluorescence due the depletion of reagents.
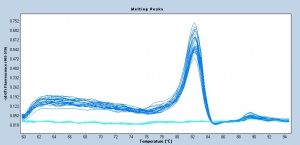
Following the RRT-PCR amplification measurements, a melt curve is completed. Melt curves assess the dissociation of double-stranded DNA while the sample is heated. As the temperature is increased, double-stranded DNA ‘melts’ as the strands dissociate. As discussed above, the fluorescent dye used in RRT-PCR associates with double-stranded DNA and fluorescence measurements will decrease as the temperature increases. In RRT-PCR, the melt curve is used to confirm that a single amplification product was generated during the reaction. If additional products were present, the melt curve would presumably show additional peaks. Why might this be true? Can you think of a scenario where two different products would produce a single peak in a melt curve?
Protocols
Part 1: Setup quantitative PCR assay
- Label 8 microcentrifuge tubes according to the samples you will prepare for RRT-PCR.
- Obtain the cDNA aliquots from the front lab bench (sample #1, #2, #3).
- Each reaction should contain 4 μL of cDNA (at 6.25 ng/uL), 2 μL of your diluted primer solution, and water for a total volume of 12.5 μL. For each sample/primer set combination, prepare enough for 2.5 reactions.
- Add your reactions to the plate on the front bench according to the map. You will add 12.5 μL to each well.
- Once everyone has added their reactions, we will add 12.5 μL of the 2X Syber Green master mix to each well.
For your reference, the PCR cycling conditions will be those shown below:
| Stage | Cycles | Details |
|---|---|---|
| 1 | 1 | 95°C for 10 min |
| 2 | 40 | 95°C for 15 sec 60°C for 30 sec 72°C for 30 sec |
| 3 | 1 | 4°C hold |
Part 2: Discussion of Research article details
Reagents
Next day: Journal club I
Previous day: Assess cell survival and harvest RNA for quantitative PCR assay