Difference between revisions of "20.109(F20):M3D2"
Noreen Lyell (Talk | contribs) (→Introduction) |
Noreen Lyell (Talk | contribs) (→Part 2: Review sgRNA expression plasmid construction strategy) |
||
| Line 21: | Line 21: | ||
===Part 2: Review sgRNA expression plasmid construction strategy=== | ===Part 2: Review sgRNA expression plasmid construction strategy=== | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
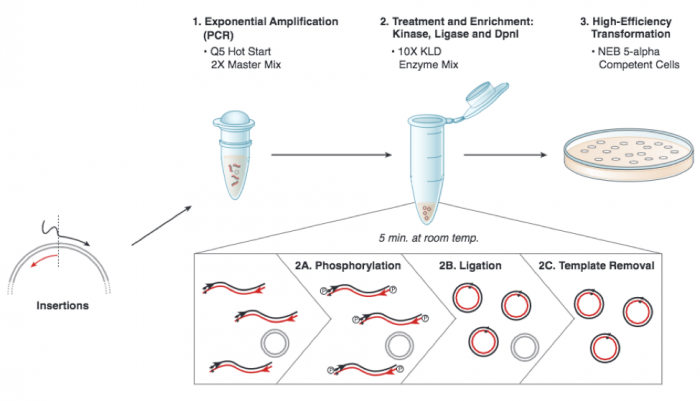
We will be using the Q5 Site Directed Mutagenesis Kit from NEB to insert the gRNA sequence into an expression vector. For this procedure you will combine the gRNA primer you designed, a universal CRISPRi primer specific to pgRNA, and the pgRNA plasmid DNA encoding. DNA polymerase will copy the plasmid using the gRNA primer to insert the target sequence you selected. Following this reaction the 'mutated' product is a linear DNA fragment. To generate circular plasmids that carry the gRNA sequence, the DNA is phosphorylated then ligated. In addition, there is still parental -- that is, non-mutant -- DNA present in your reaction product. To ensure that ''only'' the gRNA-containing plasmid is used in the next steps, the parental DNA is selectively digested using the ''DpnI'' enzyme. The underlying selective property is that ''DpnI'' only digests methylated DNA. Therefore, the synthetically made (and thus non-methylated) mutant DNA is not digested, while the parental DNA is digested due to methylation by the host bacterial strain originally used to amplify it. The resulting small linear pieces of parental DNA are simply degraded by the bacteria upon transformation, whereas the intact (due to the phosphorylation and ligation reaction) circular mutant DNA is amplified by the bacteria. | We will be using the Q5 Site Directed Mutagenesis Kit from NEB to insert the gRNA sequence into an expression vector. For this procedure you will combine the gRNA primer you designed, a universal CRISPRi primer specific to pgRNA, and the pgRNA plasmid DNA encoding. DNA polymerase will copy the plasmid using the gRNA primer to insert the target sequence you selected. Following this reaction the 'mutated' product is a linear DNA fragment. To generate circular plasmids that carry the gRNA sequence, the DNA is phosphorylated then ligated. In addition, there is still parental -- that is, non-mutant -- DNA present in your reaction product. To ensure that ''only'' the gRNA-containing plasmid is used in the next steps, the parental DNA is selectively digested using the ''DpnI'' enzyme. The underlying selective property is that ''DpnI'' only digests methylated DNA. Therefore, the synthetically made (and thus non-methylated) mutant DNA is not digested, while the parental DNA is digested due to methylation by the host bacterial strain originally used to amplify it. The resulting small linear pieces of parental DNA are simply degraded by the bacteria upon transformation, whereas the intact (due to the phosphorylation and ligation reaction) circular mutant DNA is amplified by the bacteria. | ||
Revision as of 16:41, 21 July 2020
Contents
Introduction
TECHNIQUES USED BUILD ON BASIC CONCEPTS IN PRIMER DESIGN AND PCR
Primer design
To amplify a specific sequence of DNA, you first need to design primers -- one primer that anneals at the start of the sequence of interest and a second primer that anneals at the end of the sequence of interest. Today you will design a 'forward' primer that anneals to the non-coding DNA strand and reads toward the gene that encodes TDP43 and a 'reverse' primer that anneals to the coding DNA strand at the end of the TDP43-RRM12 sequence and reads back into it. Each primer will consist of two parts: the 'landing sequence' will anneal to the sequence of interest and the 'flap sequence' will be used to add a restriction enzyme recognition sequence to your TDP43 insert.
Polymerase chain reaction (PCR)
Protocols
Part 1: Practice gRNA design principles
WRITE EXERCISE USING REFERENCE PAPER AND JOSEPHINES PRACTICE SHEET
Part 2: Review sgRNA expression plasmid construction strategy
We will be using the Q5 Site Directed Mutagenesis Kit from NEB to insert the gRNA sequence into an expression vector. For this procedure you will combine the gRNA primer you designed, a universal CRISPRi primer specific to pgRNA, and the pgRNA plasmid DNA encoding. DNA polymerase will copy the plasmid using the gRNA primer to insert the target sequence you selected. Following this reaction the 'mutated' product is a linear DNA fragment. To generate circular plasmids that carry the gRNA sequence, the DNA is phosphorylated then ligated. In addition, there is still parental -- that is, non-mutant -- DNA present in your reaction product. To ensure that only the gRNA-containing plasmid is used in the next steps, the parental DNA is selectively digested using the DpnI enzyme. The underlying selective property is that DpnI only digests methylated DNA. Therefore, the synthetically made (and thus non-methylated) mutant DNA is not digested, while the parental DNA is digested due to methylation by the host bacterial strain originally used to amplify it. The resulting small linear pieces of parental DNA are simply degraded by the bacteria upon transformation, whereas the intact (due to the phosphorylation and ligation reaction) circular mutant DNA is amplified by the bacteria.
Each group will set up one reaction, for your insertion. Meanwhile, the teaching faculty will set up a single positive control reaction, to ensure that all the reagents are working properly. You should work quickly but carefully, and keep your tube in a chilled container at all times. Please return shared reagents to the ice bucket(s) from which you took them as soon as you are done with each one.
- Get a PCR tube and label the top with your team color and lab section (write small!).
- Add 10.25 μL of nuclease-free water.
- Add 1.25 μL of your primer mix (each primer should be at a concentration of 10 μM).
- Add 1 μL of pgRNA plasmid DNA (concentration of 25 ng/μL).
- Lastly, use a filter tip to add 12.5 μL of Q5 Hot Start High-Fidelity 2X Master Mix - containing buffer, dNTPs, and polymerase - to your tube.
- Once all groups are ready, we will begin the thermocycler, under the following conditions:
| Segment | Cycles | Temperature | Time |
|---|---|---|---|
| Initial denaturation | 1 | 98 °C | 30 s |
| Amplification | 25 | 98 °C | 10 s |
| 55 °C | 30 s | ||
| 72 °C | 2 min | ||
| Final extension | 1 | 72 °C | 2 min |
| Hold | 1 | 4 °C | indefinite |
- After the cycling is completed, you will complete the KLD reaction (which stands for "kinase, ligase, DpnI") using
- 1 μL of your amplification product
- 5 μL 2X KLD Reaction Buffer
- 1 μL KLD Enzyme Mix, and
- 3 μL nuclease-free water.
- Incubate the reaction for 5 min at room temperature.
- Then, use 5 μL of the KLD reaction product to complete a transformation into an E. coli strain (NEB 5α cells of genotype fhuA2 Δ(argF-lacZ)U169 phoA glnV44 Φ80 Δ(lacZ)M15 gyrA96 recA1 relA1 endA1 thi-1 hsdR17) that will amplify the plasmid such that you are able to confirm the appropriate insertion (or 'mutation') was incorporated. The transformation procedure will be as follows:
- Add 5 μL of KLD mix to 50 μL of chemically-competent NEB 5α.
- Incubate on ice for 30 min.
- Heat shock at 42 °C for 30 s.
- Incubate on ice for 5 min.
- Add 950 μL SOC and gently shake at 37 °C for 1 h.
- Spread 50 μL onto LB+Amp plate and incubate overnight at 37 °C.
Part 3: Align gRNA sequences
Reagents list
- Q5 Site Directed Mutagenesis Kit (from NEB)
- Q5 Hot Start High-Fidelity 2X Master Mix: propriety mix of Q5 Hot Start High-Fidelity DNA Polymerase, buffer, dNTPs, and Mg2+
- 2X KLD Reaction Buffer
- 10X KLD Enzyme Mix: proprietary mix of kinase, ligase, and DpnI enzymes
- Universal CRIPSRi reverse primer 5' - ACT AGT ATT ATA CCT AGG ACT GAG CTA GC - 3'
- SOC medium: 2% tryptone, 0.5% yeast extract, 10 mM NaCl, 2.5 mM KCl, 10 mM MgCl2, 10 mM MgSO4, and 20 mM glucose
- LB+Amp plates
- Luria-Bertani (LB) broth: 1% tryptone, 0.5% yeast extract, and 1% NaCl
- Plates prepared by adding 1.5% agar and 100 μg/mL ampicillin (Amp) to LB
Next day: Analyze ethanol yield data