Difference between revisions of "Assignment 9, Part 3: Fitting your data"
Juliesutton (Talk | contribs) |
Juliesutton (Talk | contribs) |
||
| Line 12: | Line 12: | ||
#Record your estimates for &Delta H and &Delta S. Calculate Tm. | #Record your estimates for &Delta H and &Delta S. Calculate Tm. | ||
#How do these thermodynamic parameters compare to the predicted values you obtained from DINAmelt or OligoCalc? | #How do these thermodynamic parameters compare to the predicted values you obtained from DINAmelt or OligoCalc? | ||
| + | }} | ||
| + | |||
| + | ==Residual plot== | ||
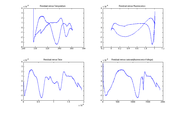
| + | [[Image:Residual plot for DNA data.png|thumb|right|Residuals plotted versus time, temperature, fluorescence, and the cumulative sum of fluorescence]] | ||
| + | Observed values differ from predicted values because of noise and systematic errors in the model. Residuals are the difference between experimental observations and model predictions, ''V<sub>f,model</sub>''−''V<sub>f,measured</sub>''. Ideally, the residuals should be random and identically distributed. | ||
| + | |||
| + | The plots at right show ''V<sub>f,model</sub>''−''V<sub>f,measured</sub>'', versus temperature, time, fluorescence, and the cumulative sum of fluorescence. The residuals are clearly not random and identically distributed. This suggests that the model does not perfectly explain the observations. The scale of the plot is much smaller than the data plot — about one percent of the data scale. | ||
| + | |||
| + | A perfect model might require dozens of added parameters and additional physical measurements. | ||
| + | |||
| + | Plotting the residuals versus different variables can help suggest what factors are not modeled well. | ||
| + | |||
| + | {{Template:Assignment Turn In|message= For one fit result, plot the residuals vs. | ||
| + | # Time, | ||
| + | # Temperature, and | ||
| + | # Fluorescence. | ||
}} | }} | ||
Revision as of 21:31, 8 November 2017
Congratulations! you should now have a working version of your analysis code.
Estimate model parameters for real data
Use your newly developed code to estimate the parameters associated with a set of DNA melting data that you took using your instrument. (You may use a data set you took from a previous week.)
Tip: you will be running this type of operation many times for many different DNA melting curves in the next couple weeks. It may be helpful to write a function to make this task easily repeatable.
Residual plot
Observed values differ from predicted values because of noise and systematic errors in the model. Residuals are the difference between experimental observations and model predictions, Vf,model−Vf,measured. Ideally, the residuals should be random and identically distributed.
The plots at right show Vf,model−Vf,measured, versus temperature, time, fluorescence, and the cumulative sum of fluorescence. The residuals are clearly not random and identically distributed. This suggests that the model does not perfectly explain the observations. The scale of the plot is much smaller than the data plot — about one percent of the data scale.
A perfect model might require dozens of added parameters and additional physical measurements.
Plotting the residuals versus different variables can help suggest what factors are not modeled well.
| |
For one fit result, plot the residuals vs.
|
Finding double stranded DNA fraction from raw data
The inverse function of the melting model with respect to Vf,measured(t) is helpful to visualize discrepancies between the model and experimental data caused by random noise in Vf,measured and systematic error in the model Vf,model. The function,
- $ C_{ds,inverse-model}(V_{f,measured}(t)) = \frac{V_{f,measured}(t) - K_{offset}} {K_{gain} S(t) Q(t)} $,
is itself a model. This model estimates the concentration of double stranded DNA based on the observations $ V_{f,measured}(t) $ and the models for bleaching and quenching.
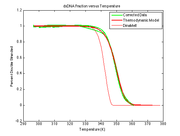
The estimated melting curve may be directly compared with simulations, measurements or other predictions of the true melting curve. The plot at right shows an example of Cds,inverse-model(t) versus Tsample(t). The estimated melting curve is shifted to the right compared to the simulated melting curve, possibly due to systematic error in the sample temperature model. The estimated melting curve also serves as a comparison to the thermodynamic model developed in DNA Melting Thermodynamics, or to any other independent measurement or model of the melting curve, i.e., the concentration of dsDNA vs sample temperature.
| |
Write a function to convert fluorescence into fraction of double stranded DNA. For at least one experimental trial, plot $ \text{DnaFraction}_{inverse-model} $ versus $ T_{sample} $ (example plot). On the same set of axes plot DnaFraction versus $ T_{sample} $ using the best-fit values of ΔH and ΔS. Finally, plot simulated dsDNA fraction vs. temperature using data from DINAmelt or another melting curve simulator. |
Comparing the known and unknown samples
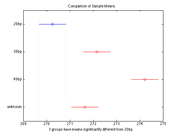
One possible way to compare the unknown sample to the three knowns is to use Matlab's anova and multcompare functions. Anova takes care of the difficulties that arise when comparing multiple sample means using Student's t-test. Read through this page: Identifying the unknown DNA sample, to learn about the statistics behind making multiple comparisons.
The following code creates a simulated set of melting temperatures for three known samples and one unknown. In the simulation, each sample was run three times. The samples have melting temperatures of 270, 272, and 274 degrees. The unknown sample has a melting temperature of 272 degrees. Random noise is added to the true mean values to generate simulated results. Try running the code with different values of noiseStandardDeviation.
% create simulated dataset
noiseStandardDeviation = 0.5;
meltingTemperature = [270 270 270 272 272 272 274 274 274 272 272 272]
+ noiseStandardDeviation * randn(1,12);
sampleName = {'20bp' '20bp' '20bp' '30bp' '30bp' '30bp' '40bp' '40bp' '40bp'
'unknown' 'unknown' 'unknown'};
% compute anova statistics
[p, table, anovaStatistics] = anova1(meltingTemperature, sampleName);
% do the multiple comparison
[comparison means plotHandle groupNames] = multcompare(anovaStatistics);
The multcompare function generates a table of confidence intervals for each possible pair-wise comparison. You can use the table to determine whether the means of two samples are significantly different. Output of multcompare is shown at right. If your data has a lot of variation, you might have to use the options to reduce the confidence level. (Or there might not be a significant difference at all.) Note that multcompare has a default confidence level of 95% (alpha = 0.05). One way to assess how confident you are in your sample identification is by finding the lowest "alpha" value needed to identify your sample.
Consider devising a more sophisticated method that uses both the ΔH° and ΔS° values, instead.
| |
|
| |
Append all of the code you wrote for Parts 1, 2 and 3 of this assignment. |
- Assignment 9 Overview
- Part 1: model function
- Part 2: test your code with simulated data
- Part 3: fitting your data
Back to 20.309 Main Page