DNA melting: Identifying the unknown sample
Overview
In the DNA lab, you will be given 4 samples containing DNA and a fluorescent dye. Samples 1, 2, and 3 are all different. The 4th sample is identical to sample 1, 2, or 3. Your job is to figure out which sample matches sample 4.
To do this, you will measure the melting temperature. Each sample has a true melting temperature $ M_j $ (where $ j $ is an integer from 1 to 4). Unknown sample 4 should have exactly the same melting temperature as sample 1, 2, or 3. Your job is to build a machine to measure the melting temperature of each sample and figure out which one matches number 4.
Procedure
Most groups measured each sample type in triplicate. (Some special students did something a little bit different.) This resulted in 12 observations, $ O_{j,i} $, where $ j $ is the sample type and $ i $ is the replicate number — an integer from 1 to 3. The majority of lab groups calculated the average melting temperature for each sample type, $ \bar{O_i} $, and guessed that sample 4 matched whichever of the known samples had the closest melting temperature.
Seems reasonable.
Except ... The observations included measurement error: $ O_{j,i}=M_j+E_{j,i} $, where $ E_{j,i} $ represents an error term. The presence of measurement error leads to the possibility that an unfortunate confluence of error terms caused you to misidentify the unknown sample. The likelihood that $ \bar{O_4} $ is closer to the mean of a sample that it does not match than the one it actually does match depends on how close the true means are together, and distribution of the error terms.
Uncertainty model
To get a handle on the possibility that your results were total crap due to bad luck alone (not incompetence), it is necessary to formulate a mathematical model for the distribution of the error terms. How about this? The error terms are normally distributed with mean $ \mu=0 $ and standard deviation $ \sigma $. (Note that the error distribution among all of the sample types is the same.) This is a simple and reasonable model. It goes without saying that the model is not perfectly correct.
Within the confines of this model, it is possible to estimate the chance that your result was a fluke.
Which unknown?
There are 6 possible pairwise hypotheses to test:
- $ M_4\stackrel{?}{=}M_1 $
- $ M_4\stackrel{?}{=}M_2 $
- $ M_4\stackrel{?}{=}M_3 $
- $ M_1\stackrel{?}{=}M_2 $
- $ M_1\stackrel{?}{=}M_3 $
- $ M_2\stackrel{?}{=}M_3 $
These are called null hypotheses. Each null hypothesis asserts that one group does not have a significantly different mean than another. Rejecting the null hypothesis means that the means are likely not the same.
Each null hypothesis can be either accepted or rejected, so there are 26=64 possible outcomes to testing all 6. In order to uniquely identify sample 4, exactly two of hypotheses 1-3 must be rejected. In other words, sample 4 must be similar to one of the other samples and dissimilar to the other two. Pretty straightforward.
Of the remaining three hypotheses, at least two of them should be rejected. The putative match for sample 4 ought be dissimilar to the other possible matches. For example, if sample 4 is the same is sample 1, it should be the case that sample 1 is distinct from sample 2 and also that sample 1 is distinct from sample 3. In this example, it doesn't matter if samples 2 and 3 are distinct or not. So the outcome of the remaining null hypotheses can go either way.
In summary, 4 of the null hypotheses must be false, 1 must be true, and 1 doesn't matter. Six of the 64 possible outcomes are consistent with a unique sample 4 identity. It is not possible to draw a conclusion from the other 58 outcomes. The following table summarizes the possibilities:
| H1 | H2 | H3 | H4 | H5 | H6 | Conclusion |
|---|---|---|---|---|---|---|
| T | F | F | F | F | X | U=1 |
| F | T | F | F | X | F | U=2 |
| F | F | T | X | F | F | U=3 |
| all others | none | |||||
Evaluating the hypotheses
Student’s t-test offers a method for assigning a quantitative measure of confidence in each null hypothesis. Essentially, the test considers the entire universe of possible outcomes of your experimental study. Imagine that you repeated the study an infinite number of times. (This may not be hard for you to imagine.) Repeating the study ad infinitum will elicit all possible outcomes of $ E_{j,i} $. The t-test categorizes each outcome into one of two realms: those that are more favorable to the null hypothesis (i.e., $ \bar{O} $ is closer to $ M $ than the result you actually got); and those that are less favorable ($ \bar{O} $ is farther from $ M $ than the result you actually got).
The t-test can be summarized by a p-value, which is equal to the percentage of possible outcomes that is less favorable to the null hypothesis than the result you got. A low p-value means that there are relatively few possible results less favorable to the null hypothesis than the one you got and many results that are more favorable. So it's probably reasonable to reject the null hypothesis.
In most circumstances, the experimenter chooses a significance level such as 10% or 5% or 1% in advance of examining the data. Another way to think of this: if you chose a significance level of 5% and repeated the study 100 times, you would expect to incorrectly reject the null hypothesis because of bad luck on 5 occasions.
Multiple comparisons
A problem arises when you use the t-test to evaluate several hypotheses. The possibility of falsely rejecting the null hypothesis for an individual test is bounded by the significance level, say 5%. If the experimental question requires 6 hypothesis tests, the overall chance of at least one false rejection will be $ 1-0.95^6\approx 26% $. The multcompare function in MATLAB implements a correction to the t-test procedure to assure that the family-wise error rate (FWER) is less than the significance level. In other words, the chance of any of the six hypotheses being incorrectly rejected due to bad luck is less than the FWER. You can set the FWER that multcompare uses via an optional argument.
If you used multcompare in your analysis, a good measure of confidence is the FWER that you chose. Since all the hypotheses may not be required to uniquely identify the unknown, the FWER is slightly conservative. So what. You required more things to be true than you strictly needed to. But it is likely that you would have gained very little by removing the unnecessary hypothesis from consideration. It is even more likely that the error terms did not perfectly satisfy the assumptions of test, so your calculation is at best an approximation of the possibility of this type of error.
Choosing the significance level
The significance level sets a bound on the chance of accidentally rejecting a null hypothesis that ought to have been accepted. Alert readers will have mused, "wait a minute ... the problem can also occur the other way around: a null hypothesis might be erroneously accepted."
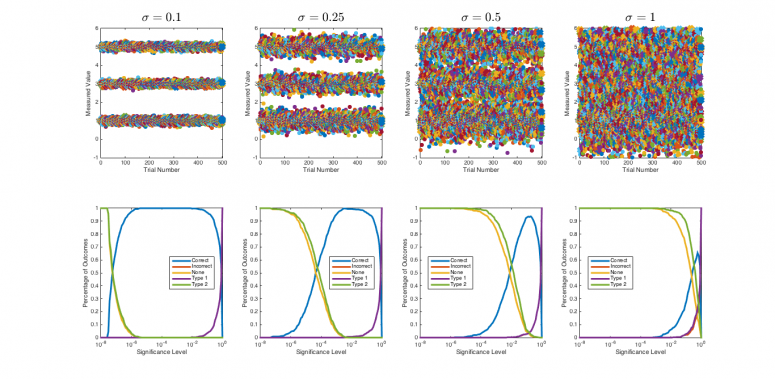
The plot below shows a simulation of using the multiple comparison test to identify an unknown sample over a range of measurement error magnitudes and significance levels. There are 4 simulated sample types with true means 1, 3, 5, and 1. (The unknown is sample 1.) The upper plots show datasets for one hundred simulated studies where each sample type was measured 3 times in the presence of various values of noise, $ \sigma $. The lower plots show significance level on the horizontal axis. For each significance level, three quantities are plotted: fraction of correct identifications (blue), incorrect identifications (red), and outcomes for which no conclusion can be drawn (gold). Making the significance threshold smaller reduces the number of erroneous identifications. But there is also a cost: a lower significance threshold increases the number of cases where no conclusion can be reached.
Unsurprisingly, less noise in the data produces better results.
The significance level can be chosen as appropriate for the circumstance. Falsely implicating a murder suspect has a very high cost, so it is probably wise to choose a low significance threshold and bear the increased risk that you might not be able to draw a conclusion at all.
What significance level to report
In the best of all cases, you would choose the significance level before you started the experiment and report that — something like p<0.01.
You can probably think of ways the simple error model this analysis relies on might be deficient. For example, there is no attempt to include any kind of systematic error. If there were significant systematic error sources in your experiment, your estimate of the likelihood of an unlucky accident may be far from the truth. Because most real experiments do not perfectly satisfy the assumptions of the test, it is frequently ridiculous to report an extremely small p-value. (That doesn't stop people from doing it, though.)
Meaning of a t-test p value
A few people argued that the p-value of a single hypothesis test is good measure of confidence. In the example above, where the unknown was sample 3, the confidence measure would be the p-value associated with hypothesis 3. Consider the two situations summarized below. In both cases, the significance level was chosen to be 5%.
- hypothesis 2 has a p-value of 4.9% (so it is rejected) and hypothesis 3 has a p-value of 5.1% (not rejected)
- hypothesis 2 has a p-value of 4.9% (rejected) and hypothesis 3 has a p-value of 90% (not rejected)
The p-value characterizes how unlikely a particular result was in the case where the null hypothesis is true. It does not compare the relative likelihood of one hypothesis to another. Clearly, there is a much smaller chance that the sample was erroneously identified in the second case above, even though the p-values for hypothesis 2 are identical. (Possibly the means of samples 2 and 3 are farther apart in the second case.)
Multiple comparisons in MATLAB
One possible way to compare the unknown sample to the three knowns is to use Matlab's anova and multcompare functions. Anova takes care of the difficulties that arise when comparing multiple sample means using Student's t-test.
The following code creates a simulated set of melting temperatures for three known samples and one unknown. In the simulation, each sample was run three times. The samples have melting temperatures of 270, 272, and 274 degrees. The unknown sample has a melting temperature of 272 degrees. Random noise is added to the true mean values to generate simulated results. Try running the code with different values of noiseStandardDeviation.
% create simulated dataset
noiseStandardDeviation = 0.5;
meltingTemperature = [270 270 270 272 272 272 274 274 274 272 272 272]
+ noiseStandardDeviation * randn(1,12);
sampleName = {'20bp' '20bp' '20bp' '30bp' '30bp' '30bp' '40bp' '40bp' '40bp'
'unknown' 'unknown' 'unknown'};
% compute anova statistics
[p, table, anovaStatistics] = anova1(meltingTemperature, sampleName);
% do the multiple comparison
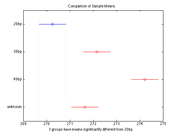
[comparison means plotHandle groupNames] = multcompare(anovaStatistics);
The multcompare function generates a table of confidence intervals for each possible pair-wise comparison and plots the result, shown at right. You can set the alpha value with an optional argument like this: [comparison means plotHandle groupNames] = multcompare(anovaStatistics, 'Alpha', 0.01));.
If you'd like to devise a more sophisticated method to analyze the data, feel free. Run your idea by an instructor before you start.