20.109(F17):Complete biochemical experiment (Day5)
Contents
Introduction
To this point in the module we have been using the CometChip assay to study DNA damage in the context of the BER pathway. Specifically, we considered the role of oxidative stress in generating base lesions. It is important to remember that the data collected using the CometChip assay is not specific to base lesions and that the DNA damage observed may be the result of various types of DNA lesions, including base excisions, abasic sites, strand breaks, and crosslinks. The experimental work you complete in the next three laboratory classes will focus on a technique used to measure DNA double-strand breaks.
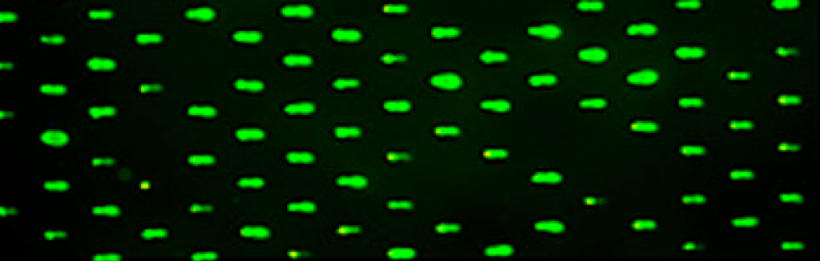
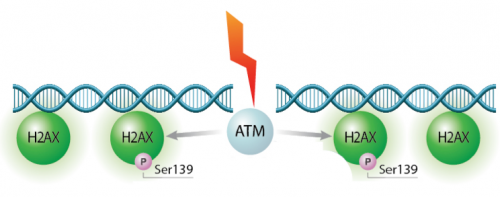
In eukaryotes, including humans, DNA is tightly wound around histone groups. H2AX is a member of the core group of histones that contributes to nucleosome formation and DNA structure. When a DNA double-strand break is introduced into the genome, the H2AX histones near the break are phosphorylated by the ATM kinase at residue Ser-139. Upon phosphorylation H2AX is referred to as gamma-H2AX. Given that only H2AX histones near the site of DNA damage are phosphorylated, γH2AX is a useful target when determining the abundance and location of double-strand breaks.In your γH2AX assay experiment, you will assess the effects of H2O2 and MMS on double-strand break abundance in the cell lines used for the CometChip assay. In your analysis you will interrogate the consistency of the results from these two approaches. Furthermore, you will craft a hypothesis concerning the repair efficiency of the cell lines and use the γH2AX assay to test your research question.
Protocols
Part 1: BE Communication Lab workshop
Our communication instructors, Dr. Sean Clarke and Dr. Prerna Bhargava, will join us today for a workshop on writing impactful abstracts and titles.
Part 2: Complete enzyme treatments for biochemical test experiments
Part 3: Separate CometChip 'tails' using gel electrophoresis
Part 4: Seed cells for sub-nuclear foci assay
Reagents list
Next day: Examine sub-nuclear foci abundance to measure DNA damage