Difference between revisions of "Assignment 10 Overview"
MAXINE JONAS (Talk | contribs) |
MAXINE JONAS (Talk | contribs) (→Using nonlinear regression to estimate parameters) |
||
| Line 92: | Line 92: | ||
}} | }} | ||
{{Template:Assignment Turn In|message=also your analysis: | {{Template:Assignment Turn In|message=also your analysis: | ||
| − | + | * Use bullet points to explain your data analysis methodology. | |
| − | + | * Document the regression model you used to analyze your data | |
| − | + | ** See [[DNA Melting: Model function and parameter estimation by nonlinear regression]] | |
| − | + | ** Explain the model parameters using bullet points or in a table. | |
| − | + | * Plot <math>V_{f,measured}</math> and <math>V_{f,model}</math> versus <math>T_{block}</math> for a typical run of each samples type. Use the smallest number of axes that clearly conveys the data. | |
| − | + | * For a typical curve, plot residuals versus time, temperature, and fluorescence, ([http://measurebiology.org/wiki/File:Residual_plot_for_DNA_data.png example plot]). | |
| − | + | * Provide a table of the best-fit model parameters and confidence intervals for each experimental run. Also include the estimated melting temperature for each run. | |
| − | + | * For at least one experimental trial, plot <math>\text{DnaFraction}_{inverse-model}</math> versus <math>T_{sample}</math> ([http://measurebiology.org/wiki/File:Inverse_cuvrve.png example plot]). On the same set of axes plot DnaFraction versus <math>T_{sample}</math> using the best-fit values of ΔH and ΔS. Finally, plot simulated dsDNA fraction vs. temperature using data from DINAmelt or another melting curve simulator. | |
}} | }} | ||
Revision as of 18:31, 1 November 2017
Assignment 10
Data acquisition
Identify unknown sample
You will receive 1.5 mL each of four samples. Three of the samples will be identified by their sequence, salt ion concentration, and degree of complementarity (see these DNA_Melting:_DNA_Sequences). The fourth sample matches one of the three identified samples. You will not be told which one.
- Acquire melting curves for the known and unknown samples.
- You may want to run some or all of the samples more than once to provide more confidence in your result.
- Identify the unknown sample and report your confidence in the result.
- See Identifying the unknown DNA sample for some ideas on identifying the sample.
- You may use a different statistical procedure, if you like. Be sure to document the procedure you used.
Analysis
We set out to measure the fraction of dsDNA versus temperature. As discussed in lecture, factors like photobleaching, thermal quenching, and the difference between the block and sample temperature distorted the measurement in various ways. For the final report, you will carefully analyze your data using a mechanistic model to factor out the melting dynamics from the shortcomings of the measurement technique.
In an ideal world, you would have your analysis code written before the data is collected. In the real world, data collection involves a lot of waiting around, and it might be a good use of your time to work on the analysis code and collect data in parallel. In that case, it is still important to ensure that your data makes sense as it comes in. For example, is the trend in the melting temperature what you expected? … e.g. did longer oligos have a higher melting temperature than shorter ones? The following section provides a quick way to estimate melting temperatures from the raw data.
Estimating melting temperatures from raw data
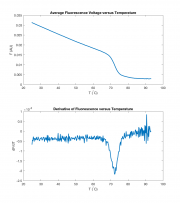
A frequently-used, quick-and-dirty proxy for melting temperature is the peak value of the fluorescence voltage's derivative as a function of temperature. The peak of the derivative is the temperature at which the equilibrium is changing most rapidly. It's not exactly the same as the melting temperature, but it allows for quick comparison of results.
You can't just take the derivative of the raw signal. The raw data is not guaranteed to be a function at all. One way to take a good derivative is to combine the fluorescence voltage values that fall into a particular range of temperatures and then take the average of all those values — a process sometimes called binning or discretization. Temperature bins of about a quarter of a degree work well. The code below demonstrates how to accomplish this in MATLAB. Because of the difference between the block and sample temperatures, it is best to do this for just the heating or melting portion of the curve.
Assuming the variables temperature and fluorescence are defined, the following Matlab code fragment below computes ΔF/ΔT for the heating portion of the curve. If your melting curves are very noisy, you may have to adjust the code.
discretizationInterval = 0.25;
discreteTemperatureAxis = 20: discretizationInterval:100; % center value for each bin
discreteTemperaturBinEdges = [ discreteTemperatureAxis - discretizationInterval / 2,
discreteTemperatureAxis(end) + discretizationInterval / 2 ];
temperatureBinIndex = discretize( temperature, discreteTemperaturBinEdges );
binnedFluorescence = accumarray( temperatureBinIndex', fluorescence',
size( discreteTemperatureAxis' ), @mean )';
badOnes = binnedFluorescence == 0;
discreteTemperatureAxis( badOnes ) = [];
binnedFluorescence( badOnes ) = [];
figure
subplot(211)
plot( discreteTemperatureAxis, binnedFluorescence )
title( 'Average Fluorescence Voltage versus Temperature')
xlabel( 'T (^{\circ}C)' )
ylabel( 'F (AU)' )
dFluorescenceDTemperature = diff( binnedFluorescence ) ./ diff( discreteTemperatureAxis );
derivativeAxis = mean( [ discreteTemperatureAxis(1:(end-1)); discreteTemperatureAxis(2:end) ] );
subplot(212)
plot( derivativeAxis, dFluorescenceDTemperature);
title( 'Derivative of Fluorescence versus Temperature' )
xlabel( 'T (^{\circ}C)' )
ylabel( 'dF/dT' )
Using nonlinear regression to estimate parameters
The DNA Melting: Model function and parameter estimation by nonlinear regression wiki page will guide you in writing the model function used to estimate the relevant DNA melting parameters.
| |
Turn in the opening of a lab report:
|
| |
also your analysis:
|
Resources
Background reading
- Electronics primer
- Real electronics
- Impedance analysis
- Transfer functions and Bode plots
- Input and output impedance
- DNA melting: Identifying the unknown sample
- DNA Melting Thermodynamics
- DNA Melting: DNA Sequences
- DINAMelt Web Server
Code examples and simulations
- DNA Melting: Simulating DNA Melting - Basics
- DNA Melting: Simulating DNA Melting - Intermediate Topics
- DNA Melting: Model function and parameter estimation by nonlinear regression
Subset of datasheets
(Many more can be found online or on the course share)
- Overview
- Part 1: convolution Let's check that you're comfortable with convolution and the lock-in amplifier design;
- Part 2: lock-in amplifier
Back to 20.309 Main Page