Spring 2012: Jonathan Gootenberg Lab 2
From Course Wiki
Contents
Lab 2: Fluorescence Microscopy
UNDERGOING EDITS
Goals
- Controlling the microstepper with the DAQ/Matlab interface
- Controlling the camera with
- Learning more about 3d deconvolution
- Test how it deconvolution is improved with actual PSF vs. theoretical PSF of the microscope
Methodology
Materials
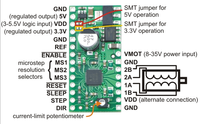
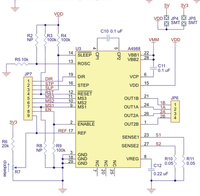
- Pololu A4983 Stepper Motor Driver Carrier with Voltage Regulators[1]
- Thorlabs 8 mm Travel Stepper Motor Actuator[2]
- Matlab
- Matlab Image Acquisition Toolbox[3]
- Matlab Allied Vision Technologies Adaptor[4]
- National Instruments USB Data Acquisition[5]
- Epi-fluorescence Microscope[6] with parts from Thorlabs
- Submicron beads, small enough to be considered point sources
- ImageJ[7]
- ImageJ plugins: Deconvolution Lab[8] and Diffraction PSF 3D[9]. Manually save in the plugins folder in ImageJ.
Deconvolution Methods for 3D Fluorescence Microscopy Images[10]
Microscope construction
- The microscope was a reanimation of the teaching microscope from the 20.309 lab. We changed the lenses a bit.
- The scope contains a 8.33x beam expander and 40x objective as of now. The objectives are changeable, but require some adjustment of the 200mm tube lens.
Laser Alignment
- The neutral density filter ensures we do not burn our eyeballs while aligning the laser. 532 nm protective goggles should be worn when appropriate.
- T-shirt targets were used as guides to center the expanded beam with the mirrors.
- With barrier filter still in place (to limit slide photobleaching), we used a fluorescent reference slide and Steve's Super Special Software (or something like that) called Capture To Memory to look at the intensity histogram and redirect the mirror just prior to the CCD camera to maximize light homogeneity.
- Profit.
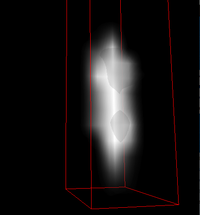
PSF Scanning
- Open GUI
- Turn on sleep, manually focus on psf sub-micron fluorescent beads (190 nm)
- Turn off sleep, run psf scan on GUI
- Take image stack, open imageJ with deconvolution plugin
Creating a theoretical PSF
- Download Diffraction PSF 3D to the plugin folder in ImageJ
- Open ImageJ
- Under plugins select Diffraction PSF 3D
- Enter parameters for
- a) Index of Refraction of the media = 1.398
- b) Numerical Apeture n*sin(theta) = 0.65
- c) Wavelength = Nile Red (525-605) – 590 because the cut off of the barrier filter is 590nm
- d) Longditudinal Spherical Aberration = 0
- e) Image pixel spacing = 7.4um/40x = 185nm
- f) Slice spacing = 100nm
- g) Width = 20px
- h. Height =20px
- i) Depth = 84px
- Ctrl + E, change first box to 10.0. Enter.
- Image > Type > 8-bit or 8-bit color
- Press Ctrl + Shift + C to change intensity thresholds and contrast. Apply.
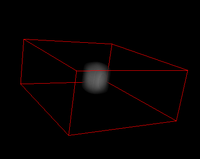
- Use Plugins > 3D to visualize
Results
Code
Code can be viewed in my Public until ~June 2012. Afterwards, it can be found on the courseshare under Students > Labs > PSF > Spring 2012.
PSFconstruct.m
- Takes in:
- A 3D data stack of the original files of type double
- xbounds and ybounds, two vectors which define the ROI ([min max])
- Defines xSize, ySize, zSize as original dimensions of new stack
- Iterate through z-slices, write a new .tif file with desired dimensions defined by xbounds and ybounds
PSFmicrometer.m
- Establishes digital channel in 'Dev1'
- Adds 6 digital output lines [Direction Step M1 M2 M3 _Sleep_] [1]
- 0 Clockwise, 1 Counterclockwise
- Driver will step (theoretically) in the middle of the rise of the Step voltage from low to high
- _Sleep_ is 0 sleep, 1 no sleep
- M1 M2 M3 determine step size
| MS1 | MS2 | MS3 | Microstep Resolution |
|---|---|---|---|
| Low | Low | Low | Full Step |
| High | Low | Low | Half Step |
| Low | High | Low | Quarter Step |
| High | High | Low | Eighth Step |
| High | High | High | Sixteenth Step |
- In the future, user can set the zSampleDistance (aka the zdepth of the scan) and nyquistStepDistance
- Takes these values and determines which step size to use in microns and the value to convert to binary to send to the Driver in M1 M2 and M3
- Actual step size will always be the step smaller than the nyquistStepDistance
- Determines numberOfSteps which is just zdistance/stepdistance
- Moves Stage
- Half the zdistance back, quickly
- Full zdistance scan forward through the area of interest, slowly taking photos at each stop
- Half the zdistance back to original place
- Stops the channel
Cite error: <ref> tags exist, but no <references/> tag was found