20.109(S22):M2D5
Contents
Introduction
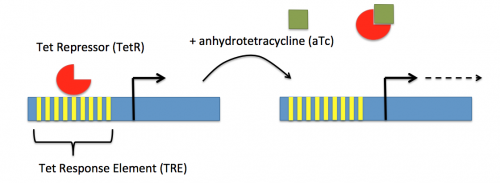
In contrast, expression of the gene encoding dCas9 within pdCas9 is regulated by an inducible promoter (pLtetO-1). An inducible promoter is 'off' unless the appropriate molecule is present to relieve repression. In the case of the CRISPRi system, expression of the gene encoding dCas9 is inhibited due to the use of a tet-based promoter construct. Tet is shorthand for tetracycline, which is an antibiotic that inhibits protein synthesis through preventing the association between charged aminoacyl-tRNA molecules and the A site of ribosomes. Bacterial cells that carry the tet resistance cassette are able to survive exposure to tetracycline by expressing genes that encode an efflux pump that 'flushes' the antibiotic from the bacterial cell. To conserve energy, the tet system is only expressed in the presence of tetracyline. In the absence of tetracycline, a transcription repressor protein (TetR) is bound to the promoter upstream of the tet resistance cassette genes. When tetracycline is present, the molecule binds to TetR causing a confirmational change that results in TetR 'falling off' of the promoter. In the CRISPRi system, the tet-based promoter construct upstream of the gene that encodes dCas9 is 'off' unless anhydrotetracyline (aTc), an analog of tetracyline, is added to the culture media. Why is it important to use an analog rather than the actual antibiotic?Taken together, the sgRNA_target is constitutively transcribed and, thereby, always present. The dCas9 protein is only present when aTc is added. Thus, gene expression is only altered when aTc is present. In this, when dCas9 is expressed it forms a complex with the sgRNA_target. The sgRNA target then 'seeks out' the target within the host genome. When the targeted sequence is recognized, the complex binds and acts as a 'roadblock' by prohibiting RNAP access to the sequence. Because the targeted gene is not able to be transcribed, the protein encoded by that gene is not synthesized. In our experiments, we hypothesize that the absence of specific proteins, or enzymes, involved in anaerobic fermentative metabolism will increase the yield of ethanol.
Protocols
Part 1: Examine psgRNA_target sequencing results
Your goal today is to analyze the sequencing data for you two potential mutant psgRNA clones - two independent colonies from your cloning reaction - and then decide which colony to proceed with for the CRISPRi engineering of the E. coli MG1655 fermentation pathway.
Retrieve psgRNA_target sequence results from Genewiz
- Your sequencing data is available from Genewiz. For easier access, the information was uploaded to the [20.109(S22):Class_data Class Data tab].
- Download the zip folder with your team sequencing results and confirm that there are 8 files saved in the folder.
- For each sequencing reaction, you should have one .abi file and one .seq file.
- Open one of the .abi files.
- This file contains the chromatogram for your sequencing reaction. Scroll through the sequence and ensure that the peaks are clearly defined and evenly spaced. Low signal (or peaks) or stacked peaks can provide incorrect base assignments in the sequence.
- Open one of the .seq files.
- This file contains the base assignments for your sequencing reaction. The bases are assigned by the software from the chromatogram sequence.
- The start of the a sequencing reaction result often contains several Ns, which indicates that the software was unable to assign a basepair.
In your laboratory notebook, complete the following:
- Given the chromatogram result, why might the software assign Ns in the start of the sequence?
- Visually inspect the chromatograms for all of your sequencing results.
- Do the peaks appear clearly defined or is there overlap? What might this indicate about the quality of your sequencing results?
- Do the peaks extend above the background signal? What might this indicate about the quality of your sequencing results?
Confirm sgRNA_target sequence in psgRNA_target using SnapGene
You should align your sequencing data with a known sequence, in this case the gRNA target sequence you selected, to identify any unintended base changes that may have occurred. There are several web-based programs for aligning sequences and still more programs that can be purchased. The steps for using SnapGene are below. Please feel free to use any program with which you are familiar.
- Generate a new DNA file that contains the sgRNA oligo you designed on M2D2. This file should contain only the target sequence you selected and the dCas9 tag sequence.
- Generate an additional new DNA file that contains the results from the sequencing reaction completed by Genewiz.
- For each sequencing result you should generate a distinct new DNA file. Remember you should have a forward and reverse sequencing result for each of your clones!
- Paste the sequence text from your sequencing run into the new DNA file window. If there were ambiguous areas of your sequencing results, these will be listed as "N" rather than "A" "T" "G" or "C" and it's fine to include Ns in the query.
- The start and end of your sequencing may have several Ns. In this case it is best to omit these Ns by pasting only the 'good' sequence that is flanked by the ambiguous sequence.
- To confirm the sgRNA sequence in your clones, open one of the forward sequencing results files generated in the previous step.
- Select 'Tools' --> 'Align to Reference DNA Sequence...' --> 'Align Full Sequences...' from the toolbar.
- In the window, select the file that contains the sgRNA oligo sequence and click 'Open'.
- A new window will open with the alignment of the two sequences. The top line of sequence shows the results of the sequencing reaction and the bottom line shows the oligo you designed.
- Are there any discrepancies or differences between the two sequences? Scroll through the entire alignment to check the full sequencing result and note any basepair changes.
- Follow the above steps to examine all of your sequencing results. Remember: you used a forward and a reverse primer to interrogate both potential psgRNA_target plasmids.
- From the alignments, determine which psgRNA_target has the correct sgRNA_target sequence.
- If both clones contain the correct sequence choose either co-transformant to use to test ethanol / acetate yield. If only one is correct, then this is the co-transformant you will use. If neither of your plasmids carry the appropriate insert, talk to your Instructor.
In your laboratory notebook, complete the following:
- Attach a screenshot for each alignment.
- Record which clone contains the correct sgRNA_target sequence.
Part 2: Prepare media for fermentation product yield experiment
To test the effect of sgRNA_target on increasing the production of ethanol / acetate, the co-transformed E. coli MG1655 cells will be incubated both with and without oxygen. Remember from lecture that cells grown anaerobically should produce more fermentation products; however, the goal of using the CRISPRi system is to further enhance the production of ethanol / acetate by manipulating gene expression of an enzyme in the anaerobic fermentative pathway.
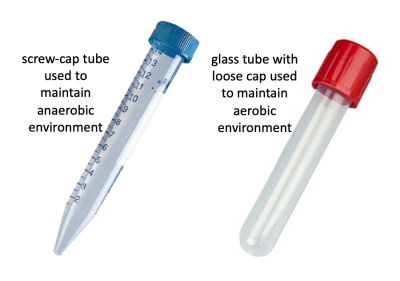
Before we look at the specific conditions that were tested, let's first review how anaerobic and aerobic cultures will be maintained. Rather than using anaerobic chambers or gas replacement, a much simpler technique was employed to omit oxygen from the cultures for this experiment. Because screw-cap tubes maintain a tight seal, cultures can be maintained in a low-O2 environment. For this technique cultures are grown at least 24 hrs to ensure the following: 1. that the oxygen present at the time of inoculation is depleted by the growing cells and 2. that the cells grow in the O2-depleted environment long enough to undergo anaerobic fermentative metabolism. Aerobic cultures were maintained using standard glass test tubes with loose-fitting caps. Both culture tubes are shown in the image to the right.- Acquire 4 glass culture tubes and 4 15 mL conical tubes from the front laboratory bench and label as shown below:
- MG1655 +O2 -aTc (glass tube)
- MG1655 -O2 -aTc (15 mL conical tube)
- MG1655 +O2 +aTc (glass tube)
- MG1655 -O2 +aTc (15 mL conical tube)
- MG1655 +CRISPRi +O2 -aTc (glass tube)
- MG1655 +CRISPRi -O2 -aTc (15 mL conical tube)
- MG1655 +CRISPRi +O2 +aTc (glass tube)
- MG1655 +CRISPRi -O2 +aTc (15 mL conical tube)
- Be sure to include your team information on each tube!
- Obtain a bottle of LB broth from the front laboratory bench.
- Transfer 5 mL of the media into each tube labeled for inoculation with MG1655 (alone, not co-transformed).
- Calculate the volume of the chloramphenicol and ampicillin antibiotic stocks that are required to maintain the pdCas9 and pgRNA plasmids, respectively, in the remaining volume of the LB media using the information below:
- For chloramphenicol: the stock concentration is 25 mg/mL and the final concentration should be 25 μg/mL.
- For ampicillin: the stock concentration is 100 mg/mL and the final concentration should be 100 μg/mL.
- Add the appropriate volume of each antibiotic stock to the LB aliquot and mix thoroughly.
- Transfer 5 mL of the media you prepared in Step #5 into each tube labeled for inoculation with MG1655 +CRISPRi.
- Return your labeled tubes with media added to the front laboratory bench.
- Acquire 1 additional glass culture tube and label with team information.
- Transfer 5 mL of the media containing chloramphenicol and ampicillin into the glass culture tube.
- Inoculate one of the co-transformant colonies from your agar plate into the glass culture tube to prepare a starter culture.
- To inoculate the colony, use an innoculation loop to lightly scrape one colony from the plate and then swirl the loop in the media within the culture tube.
- Cap the glass culture tube and place in the turning wheel in the 37 °C incubator.
The Instructors will inoculate MG1655 or your co-transformant from the starter culture (which you prepared in Step #11) into your culture tubes (using a 1:1000 dilution of your starter culture) and add 2 μM aTc (final concentration) to the appropriate tubes. All cultures will then be incubated at 37 °C for at least 24 hrs in preparation for your next laboratory session.
In your laboratory notebook, complete the following:
- For each of the conditions that will be tested, answer the following questions:
- Is the cell culture using anaerobic fermentative metabolism?
- Do the cells contain the CRISPRi system?
- Is the CRISPRi system active (are the components expressed)?
- Which culture conditions provide controls? What does each control indicate / validate?
- In which culture condition do you expect to see the lowest yield of ethanol / acetate? Why?
- In which culture condition do you expect to see the highest yield of ethanol / acetate? Why?
Reagents list
- Luria-Bertani (LB) broth: 1% tryptone, 0.5% yeast extract, and 1% NaCl
- chloramphenicol antibiotic (from Sigma)
- ampicillin antibiotic (from Sigma)
- anhydrotetracycline (aTc) (from Sigma)
Next day: Complete CRISPRi experiment and measure fermentation products