20.109(S16):Data analysis (Day9)
Contents
Introduction
Today is the final laboratory day of Module 2 and the goal is for you to have all of your NHEJ calculations and statistical analyses complete before you leave...you may even have time to outline your results section with the data/figures you will use to craft your story for the System engineering research article.
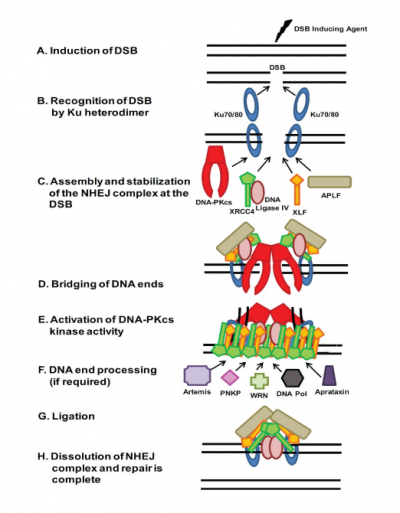
Our lecture and prelab discussions of NHEJ have primarily focused on a few key players: DNA-PKcs, Ku70, and Ku80. However, as has been noted by Prof. Samson, and as is shown in the image to the right, there are several other proteins that are involved in the process. In your experiments, you specifically assessed the role of DNA-PKcs in regard to repairing specific types of DNA damage. The review article by Davis et al. highlights the role of DNA-PK (and may be helpful as you write your research article). As a brief review, Ku80, with its dimer partner Ku70, completes the first step of NHEJ repair by binding to DNA double-strand breaks. The Ku 80/70 dimer quickly recruits DNA-PKcs, the catalytic subunit of DNA-dependent protein kinase, to form the DNA-PK complex. Although DNA-PKcs has some binding affinity for DSBs, this affinity is increased by two orders of magnitude by the presence of the Ku dimer, and the kinase activity itself cannot proceed in the absence of Ku. Interestingly, although the kinase activity of DNA-PK is known to be important, it is not certain precisely which phosphorylation events are absolutely required for NHEJ. Keep this in mind as you interpret your data and begin to consider the implications of your results.The final experimental work that you will do for this module is to complete your NHEJ inhibitor kill curve assay. Clonogenic assays of mammalian cells have been used for over 50 years, as mentioned in the methods paper by Franken et al. These experiments are useful in assessing the reproductive capacity of cells after irradiation and following other types of damage. We will diverge somewhat from the protocol within the reference paper, but it is useful for introducing terms such as the plating efficiency and the surviving fraction. Specifically, we will not fix our cells in an independent step as this does not appear to be crucial for short-term staining. Second, we will not use the crystal violet stain, which binds DNA, but instead a Coomassie derivative, which targets proteins. In fact, you may remember Coomassie as the stain used for SDS-PAGE. Protein binding by the dye occurs primarily via arginine, as well as other basic and aromatic residues, as described here. We will use a variant of the original Coomassie Brilliant Blue stain called BioSafe Coomassie.
Protocols
Part 1: Paper discussion with Prof. Leona Samson
See M2D7 for guidelines on this rescheduled discussion of the research article by Dietlein et al.
Part 2: Stain irradiated cell colonies
Your irradiated cells have now been growing for 12 days. Time to count colonies!
- Briefly observe your irradiated cells on the TC microscopes. Are you able to find some colonies? About how many cells are in some of these small dispersed clusters?
- Take your plate to the main lab for the remaining steps. First, aspirate the media. You don't need to change the yellow tip between samples if you move from highest drug dose to lowest, as that should be the order of least to most colonies.
- Rinse each well with about 2 mL of PBS.
- After removing the PBS, use a serological pipet to consistently add 2 mL of Coomassie to each well.
- Place your plate on the shaker at 80 rpm for about 1 hour.
- Use this time to complete the statistics exercise in Part 3...and perhaps begin analyzing your NHEJ efficiency data from M2D7.
- Repeat the PBS rinse.
- Let the well plate dry for a short time after aspirating the PBS.
- If you wait a very long time the stain will begin to fade.
- Finally, count the colonies in each well and document these on today's Discussion page.
- Do your best to apply a consistent standard for threshold colony size and threshold staining intensity. As long as you are consistent, there is not one right answer as to what constitutes a colony.
- One of your instructors finds it easiest to count colonies by making a dot with a lab marker as she counts each colony, right on the underside of the 6-well plate at the colony location, and then writing down each decade (10, 20, etc.) on the plate as she reaches that number so she doesn't lose her place.
- Another one likes taking a picture of her plate (using the zoom of the gel documentation station), printing it, and counting colonies on paper.
- Whether today or on your own, you should plot your data by surviving fraction for the Module 2 report. If you are feeling ambitious (and your data is appropriate) you can estimate the IC50 for your inhibitor. Does it agree with the published values?
Part 3: Applying statistical analysis to data
You may find averages, measures of spread (standard deviation or 95% confidence intervals), and t-tests useful when you report on class results.
You can practice the steps below using the highlighted cells of the for statistics tab in the Instructors Flow Cytometry Data Analysis spreadsheet. They recapitulate the normalized EGFP expression for all untreated M059K and M059J fibroblasts comprised in the instructors' M2 flow cytometry data set.
- Begin by downloading the following Excel file as a framework to carry out the basic statistical manipulations we discussed in lecture. The file is modified from one originally written by Professor Bevin Engelward.
- Find and plot 95% confidence intervals for the M059K vs. M059J normalized green signal.
- Compare the means of these two populations. At what confidence level (if any) are they different?
- Would a one-tailed or two-tailed test be more appropriate for this comparison?
When are you comfortable with the statistics tools that were presented in lecture and included in the above worksheet, use this knowledge to assess your NHEJ efficiency data.
- Also check out the other two tabs of the Instructors Flow Cytometry Data Analysis spreadsheet; they contain all the numbers you might want to compare and represent in your M2 Systems Engineering research article.
Reagent list
- PBS
- Bio-Safe Coomassie Stain (Bio-Rad)
- Mostly your brains!
Next day: Module 3 starts: Growth of phage nanowires