Difference between revisions of "20.109(S21):M1D1"
(→Part 1: Investigate our scFv clone) |
(→Part 2: Mutagensis of scFv clone using Error-prone PCR) |
||
| Line 26: | Line 26: | ||
We will use SnapGene software for our DNA visualization and alignments. Feel free to use any software you are familiar with to carry out analysis. If you are off campus you must log into a VPN connection prior to opening the SnapGene application. Here is the link to the [https://ist.mit.edu/cisco-anyconnect VPN download] and [http://kb.mit.edu/confluence/x/6QPn installation instructions]. You may need to update the MIT SnapGene license number if you have not opened the application recently. The new license information can be found [http://downloads.mit.edu/released/snapgene/group-name_registration-code.txt here]. | We will use SnapGene software for our DNA visualization and alignments. Feel free to use any software you are familiar with to carry out analysis. If you are off campus you must log into a VPN connection prior to opening the SnapGene application. Here is the link to the [https://ist.mit.edu/cisco-anyconnect VPN download] and [http://kb.mit.edu/confluence/x/6QPn installation instructions]. You may need to update the MIT SnapGene license number if you have not opened the application recently. The new license information can be found [http://downloads.mit.edu/released/snapgene/group-name_registration-code.txt here]. | ||
| − | ===Part 2: | + | ===Part 2: Mutagenesis of scFv clone using Error-prone PCR=== |
| + | To introduce random DNA mutation into our scFv clone we will carry out error-prone polymerase chain reaction (PCR). We will use special bases in the reaction that mimic oxidative damage to the DNA base. | ||
| + | <center> | ||
| + | {| border="1" | ||
| + | | | ||
| + | !component | ||
| + | |buffer | ||
| + | |MgCl2 | ||
| + | |Forward primer | ||
| + | |Reverse primer | ||
| + | |- | ||
| + | ! Final concentration | ||
| + | |10X | ||
| + | | | ||
| + | | | ||
| + | | | ||
| + | |- | ||
| + | !Volume | ||
| + | | 1X | ||
| + | | | ||
| + | | | ||
| + | |- | ||
===Part 3: Gel purify PCR products=== | ===Part 3: Gel purify PCR products=== | ||
Revision as of 18:18, 27 January 2021
Contents
Introduction: Generate single-chain antibody fragment (scFv) library
To engineer a protein-protein interaction (antibody-antigen), we will use a method called directed evolution. Directed evolution is defined as an iterative process of mutating a gene, then screening those mutants for a particular functionality, collecting the best mutants and repeating. It is a method that is inspired by the process of natural selection.
In the Wittrup lab protocol, successive rounds of random mutagenesis of the scFv gene, followed by expression of the scFv on the surface of yeast then selection for "good binders" by fluorescence activated cell sorting is carried out. This process will produce an antibody fragment with increased affinity or stability to its antigen.
Today we will walk through the technical details of random DNA mutagenesis via error-prone PCR. This process creates a pool of scFv genes that have random mutations throughout the DNA sequence. This collection of varied scFV genes are called a library.
Finally, we will outline how one integrates this library into an expression vector, also referred to as the backbone or plasmid, that can then be transformed into yeast and expressed on the surface of the cells. This method is called cloning. Integral tools when cloning DNA plasmids are restriction endonucleases.
Restriction enzyme digest
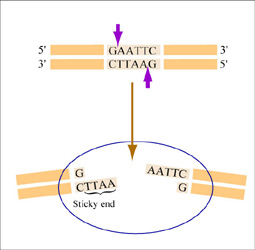
Restriction endonucleases, also called restriction enzymes, 'cut' or 'digest' DNA at specific sequences of bases. The restriction enzymes are named according to the prokaryotic organism from which they were isolated. For example, the restriction endonuclease EcoRI (pronounced “echo-are-one”) was originally isolated from E. coli giving it the “Eco” part of the name. “RI” indicates the particular version on the E. coli strain (RY13) and the fact that it was the first restriction enzyme isolated from this strain.
The sequence of DNA that is bound and cleaved by an endonuclease is called the recognition sequence or restriction site. These sequences are usually four or six base pairs long and palindromic, that is, they read the same 5’ to 3’ on the top and bottom strand of DNA. For example, the recognition sequence for EcoRI is 5’ GAATTC 3’ (see figure at right). EcoRI cleaves the phosphate backbone of DNA between the G and A of the recognition sequence, which generates overhangs or 'sticky ends' of double-stranded DNA.
Unlike EcoRI, some other restriction enzymes cut precisely in the middle of the palindromic DNA sequence, thus leaving no overhangs after digestion. The single-stranded overhangs resulting from DNA digestion by enzymes such as EcoRI are called sticky ends, while double-stranded ends resulting from digestion by enzymes such as HaeIII are called blunt ends. HaeIII recognizes 5’ GGCC 3’ and upon recognition cuts in the center of the sequence.
Protocols:
Part 1: Investigate our scFv clone
We will use SnapGene software for our DNA visualization and alignments. Feel free to use any software you are familiar with to carry out analysis. If you are off campus you must log into a VPN connection prior to opening the SnapGene application. Here is the link to the VPN download and installation instructions. You may need to update the MIT SnapGene license number if you have not opened the application recently. The new license information can be found here.
Part 2: Mutagenesis of scFv clone using Error-prone PCR
To introduce random DNA mutation into our scFv clone we will carry out error-prone polymerase chain reaction (PCR). We will use special bases in the reaction that mimic oxidative damage to the DNA base.
| component | buffer | MgCl2 | Forward primer | Reverse primer | |
|---|---|---|---|---|---|
| Final concentration | 10X | ||||
| Volume | 1X |