20.109(F21):M1D2
Contents
Introduction
The goal of this module is to test the hypothesis that exposure to As inhibits the repair of H2O2-induced DNA damage in an effort to address potential public health risks associated with combined exposures to hazardous compounds. Before we discuss the specific experiments, let's review the important background information.
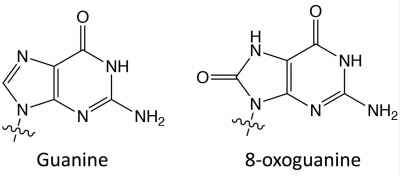
Hydrogen peroxide (H2O2) is an oxidizing agentNormal cell tissues have a basal level of DNA damage due to cell processes involved in cellular metabolism. For example, electrons can escape the electron transport chain and result in the formation of superoxide. Furthermore, defense mechanisms employed to protect the host from bacterial infection involved the release of reactive oxygen species. These reactive oxygen species are implicated in causing more than 20 types of DNA base lesions. One of the most common types of damage is the change of guanine to 8-hydroxyguanine.
Base excision repair (BER) pathway repairs damaged bases
The Base Excision Repair (BER) pathway corrects DNA damage, specifically the removal of small, non-helix distorting lesions in DNA caused by damaged bases. These lesions often result from oxidation, alkylation, deamination, and depuriniation/depryrimidination. If base lesions are not repaired, non-canonical base pairing can occur, which may result in the incorporation of an incorrect base during replication. To prevent mutations and maintain integrity of the genome, the BER pathway evolved as a highly conserved repair mechanism in both E. coli and mammals. Thus, this pathway is responsible for repairing damage before a mutation results.
The core BER pathway includes only four proteins that function to remove the damaged base and replace it with the correct base. First, a DNA glycosylase recognizes that a damaged base is present in the DNA and cleaves an N-glycosidic bond, which creates an apurinic or apyrimidinic site (referred to as an AP site in both cases). Different DNA glycosylases recognize different types of base lesions. Second, the DNA backbone is cleaved to create a single-strand DNA nick by either a DNA AP endonuclease or a DNA AP lyase. Next, a DNA polymerase incorporates the correct base using the sister strand as a template. Last, a DNA ligase completes the repair by sealing the single-strand nick, which restores integrity to the helix. For a more detailed description of the BER pathway, read this review by Robertson et al.
Arsenite (As) inhibits ligase activity
As mentioned above, the final step in BER is a ligation reaction that seals the helix. Arsenite has been shown to enhance the genotoxicity of several mutagens and inhibit ligase activity. Thus, the experiments in this module are designed to test the combined effects of exposure to H2O2, a DNA damaging agent, and As, a chemical known to decrease DNA repair capacity by inhibiting strand ligation.
Protocols
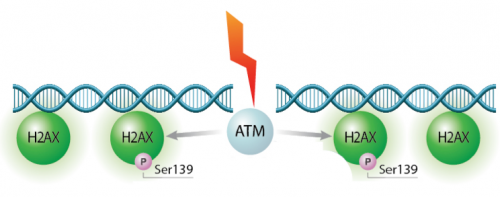
We will test the hypothesis for this module using two experiments: the γH2AX assay and the CometChip assay. Today we will start the γH2AX experiment. In eukaryotes, including humans, DNA is tightly wound around histone groups. H2AX is a member of the core group of histones that contributes to nucleosome formation and DNA structure. When a DNA double-strand break is introduced into the genome, the H2AX histones near the break are phosphorylated by the ATM kinase at residue Ser-139. Upon phosphorylation H2AX is referred to as gamma-H2AX. Given that only H2AX histones near the site of DNA damage are phosphorylated, γH2AX is a useful target when determining the abundance and location of double-strand breaks. It is important to highlight that the DNA damage expected to occur in response to H2O2 treatment is single-stranded breaks. So why are we using the γH2AX assay to measure double-stranded breaks? When DNA is damaged by multiple single-stranded breaks, double-stranded breaks can occur.
Part 1: Prepare coverslips for γH2AX assay
To ensure the steps required for preparing the coverslips for the γH2AX experiment are clear, the Instructor will provide a live demonstration of this process.
In your laboratory notebook, complete the following:
- Provide a written overview / description of the the procedure used to prepare the coverslips for cell adhesion using poly-d-lysine (from the live demonstration).
Part 2: Define treatment conditions that will be used for γH2AX experiment
A plate seeded according to the procedure demonstrated in Part 1 was used to treat cells using the H2O2 +/- As conditions detailed below.
Before you read through the protocols for the H2O2 +/- As treatments, it is important to consider what conditions were assessed in this experiment and how the experiment was designed. For the experimental question, the goal was to measure DNA damage in cells that were first exposed to As and then treated with H2O2. Given this information regarding the design of the experiment, make a list of all of the conditions that were tested with your laboratory partner. Use the following descriptions of the variables that are included in this experiment to assist you.
- Two concentrations of As will be tested: 0 μM and 2 μM
- Two concentrations of H2O2 will be tested: 0 μM and 100 μM
- Each concentration of H2O2 will be tested with each concentration of As.
In your laboratory notebook, complete the following:
- Prepare a list or table of the conditions that will be used for the γH2AX assay.
- What conditions from your list or table are controls? For what does each condition control?
- Diagram a basic experimental workflow that illustrates the order in which the treatments will be applied and when the data will be collected.
Part 3: Treat cells for γH2AX assay
For this experiment, the MCL-5 cells with be treated with H2O2 +/- As exposure. The treatments will be applied in two parts. First, cells will be incubated in As for 24 hrs to mimic exposure to the toxic metal. Second, the cells will be incubated in H2O2 for 30 min to induce DNA damage. Following treatment with H2O2 the cells will be visualized at the different timepoints to assess DNA repair.
To ensure the steps included below are clear, please watch the video tutorial linked here: [Treating cells]. The steps are detailed below so you can follow along!
Expose cells to As
Due to the timing of the As treatment, the Instructors completed the below steps using the cells that you seeded in the T75 flasks during the previous laboratory session. Using the As treated cells you will complete the steps starting at Transfer cells to coverslips.
- For each As treatment concentration, harvest and pellet cells by centrifuging at 200 g for 5 min at room temperature.
- Prepare 2 μM As treatment concentrations in media.
- Resuspend each cell pellet in 10 mL of the As-containing media.
- Transfer cell suspensions to fresh T25 flasks and incubate at 37 °C for 24 hrs.
In your laboratory notebook, complete the following:
- Calculate the amount of As stock needed to prepare the 2 μM treatment concentration.
- Stock concentration of As is 100 mM.
Transfer cells to coverslips
- Obtain As treated cell cultures from the 37 °C incubator.
- Retrieve a 12 well plate containing coverslips coated with poly-d-Lysine.
- You will need a total of two cultures treated using the following conditions: 0 μM and 2 μM As.
- Using a 10 mL pipet, transfer the cell cultures from the T25 flasks into labeled 15 mL conical tubes.
- Centrifuge the suspended cells for each condition at 500g for 5 minutes to pellet cells.
- Use pipette to remove 9.5 mls of media from conical tube (500ul with cells should remain).
- Be sure to collect all media containing As in a separate As waste container for safe disposal.
- Add 2 mL of fresh media to each cell pellet and resuspend by carefully rocking or shaking the conical tube.
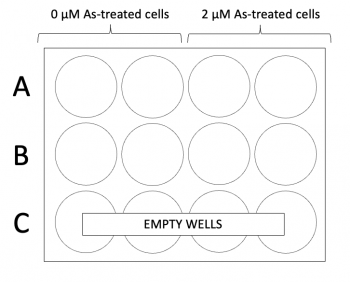
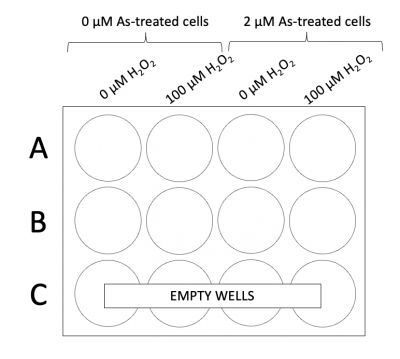
- Add 500ul of each cell suspension to the 12-well plate according to the plate map to the right. There will be approximately 25,000 cells per coverslip.
- Clearly label the lid of the 12 well plate such that the condition present in each well is evident.
- Place the plate with the cells and coverslips in the incubator for 15 minutes to allow cell adhesion.
- The remaining treatments will be performed on the cells that adhered to the coverslips.
- To prepare for the H2O2 treatment, leave your 12 well plate at 37 °C while completing the calculations for the next steps.
Treat cells with H2O2
- Prepare the 100 μM treatment concentration of H2O2 in serum-free media.
- Be sure to prepare with serum-free media as the diluent because the serum in complete culture media will inactivate H2O2.
- For the H2O2 treatment conditions, As should also be included at the concentrations previously used for the exposure step.
- Retrieve your 12-well plate from the 37 °C and carefully use a P1000 pipet to remove the liquid from each well.
- Collect the in the As waste container.
- Add 1 mL of either media or media + 100 μM H2O2 to the appropriate wells according to the plate map to the right.
- Carefully transport your 12-well plate to the 4 °C cooler and incubate for 30 min.
- Retrieve the plate from the cooler.
- Complete the steps described below under 'Fix cells for γH2AX staining'.
In your laboratory notebook, complete the following:
- Calculate the dilution of H2O2 needed to have a final concentration of 100 μM in 5ml of media.
- Stock concentration of H2O2 is 10 M and working dilution created from stock is 100 mM.
Fix cells for γH2AX staining
- Aspirate the liquid from the well and immediately add 400 μL of 4% paraformaldehyde to fix the cells.
- Incubate at room temperature for 10 min.
- Collect the 4% paraformaldehyde in the correct waste stream using a P1000 pipet.
- Wash with 500 μL of 1X PBS.
- Add 1X PBS then remove using a P1000 pipet. Collect the PBS in the correct waste stream.
- Complete a total of 2 times. After the final wash leave 1 mL of 1X PBS on the cells in the final wash.
- Leave all wells with 1 mL 1X PBS, parafilm the sides and move the 12-well plate into the 4 °C cooler.
Reagents list
- 0.1 mg/ml poly-d-Lysine solution (from Sigma)
- 4% paraformaldehyde (from VWR)
- arsenite (As) (from Sigma)
- hydrogen peroxide (H2O2) (from Sigma)
- phosphate saline buffer (PBS) (from VWR)
Next day: Use immunoflourescence staining to assess repair foci experiment