20.109(F17):Develop experiment to optimize loading variables (Day2)
Contents
Introduction
Today you will load cells into the CometChip you prepared during the previous class to test cell loading conditions in an effort to optimize the assay developed by the Engelward Laboratory for our experimental purposes.
Cell loading refers to the process by which the cells are added to the CometChip. Briefly, you will add media containing cells over the agarose-based CometChip, and gravity will pull the cells down and 'load' them into the CometChip. The experimental details of this process will be the basis of the experiments you design and perform during this laboratory meeting.
Experimental design refers to the process by which the details of an experiment are organized to ensure that the data collected are appropriate and answer the correct question. In an experiment, a treatment is intentionally imposed on a sample such that the outcome(s) can be observed. You should consider these points when designing an experiment:
- Treatments should be administered in measurable levels. The level, or amount, of treatment must be conserved across samples to limit unintended variability in the results.
- Controls should be included. The controls, or untreated samples, are a baseline to which the treated samples are compared.
- Each experiment should have only one variable. If multiple variables (e.g. treatments, conditions, etc) are included in a single experiment, the results will be inconclusive because the outcome may be attributed to any of the variables within the experiment.
- Replicates should be included. The replicates test for technical error introduced by the researcher.
Designing a successful experiment requires time, effort, and practice. Today you will design an experiment to interrogate the best conditions for loading cells into your CometChip.
Protocols
Part 1: Design experiment to optimize CometChip loading
The experiment you design will test a variable associated with loading cells into the microwells of your CometChip. The variable in this experiment is cell number. Specifically, how many cells should be added to each well to ensure the majority of the microwells are loaded? In addition, how many cells should be loaded into each microwell?
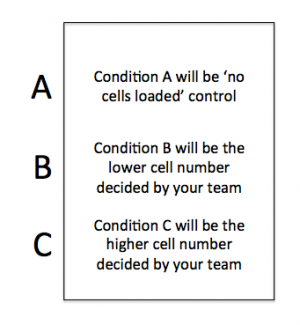
On your CometChip there is space for you to complete this experiment using three conditions. Your experiment will address the question above and the conditions will provide data that will, hopefully, answer your research question.
Experiment: Determine the number of cells needed to completely load the microwells of your CometChip
- How many cells should be loaded into each microwell?
- Consider the amount of DNA that is carried by a single mammalian cell and the detection limit provided by the SYBR gold DNA stain that will be used in your experiments. Also, use the data you collected during M1D1 Part 4 to determine how many cells can fit into a single microwell based on the dimensions provided above.
- Be sure to include any calculations or thoughts in your laboratory notebook!
- When you know how many cells you want to load into each microwell, move on to the next question.
- How many cells should be added to each well such that the desired number of cells are loaded into the microwells?
- Consider the likelihood that every cell you add to the well will fall into a microwell. Perhaps calculate the surface area of the bottom of a well (of diameter 6.35 mm) and compare this to the size of the cells as you consider this question.
- Now that you have an idea as to the number of cells that are ideal for loading, consider the conditions you will use in your experiment.
- Your team will choose two 'cell number' conditions for your experiment.
- Again, all information concerning your experimental design choices should be recorded in your laboratory notebook!
- Alert the teaching faculty when you are ready to prepare your cells for the cell loading experiment.
- Note that the tissue culture room cannot accommodate the entire class at the same time. Please be patient and complete Part 4 while you wait for an open hood.
Part 2: Prepare cells for CometChip loading
- Clean the tissue culture hood and prepare it with the supplies you will need.
- Carefully read through the protocol to ensure you have everything you need at hand.
- Retrieve the flask you seeded during the previous class session from the 37°C incubator.
- Prepare a cell suspension for use in your cell loading experiment as done previously. Briefly,
- Examine your cell culture.
- Aspirate the media.
- Wash the cells with 5 mL PBS.
- Dislodge the cells with 1 mL of trypsin.
- Incubate the cells at 37°C for 2 minutes using a timer.
- Retrieve your flask from the incubator and firmly tap the bottom to dislodge the cells.
- Check your cells using the microscope to ensure they are dislodged. They should appear round and move freely.
- Add 4 mL of media to the cells then triturate to break up cells that are clumped together and suspend them in the liquid.
- Transfer the suspended cells into a labeled 15 mL conical tube.
- Transfer 90 μL of your cell suspension from the 15 mL conical tube into a labeled eppendorf tube.
- Save the remaining cell suspension in the 15 mL conical as this is what you will use to load your CometChip!
- Clean out the tissue culture hood.
- Determine the number of cells / mL in your cell suspension as done previously. Briefly,
- Add 10 μL of trypan blue to the 90 μL aliquot of your cell suspension.
- Use the hemocytometer to count the cells, then calculate the cells / mL in your suspension.
- If necessary, use fresh media to dilute your cells such that at least 50 μL is added into each well for your determined cell number in your experiment.
Part 3: Load CometChip
You will use the MEF cells you prepared in the tissue culture hood to load the CometChip in the main laboratory. It is important that you consider the following details before entering the tissue culture room.
- The density of your MEF cell suspension will determine the volumes you load in the CometChip for Conditions B and C.
- Calculate the volume of this suspension that you will need to load for Conditions B and C. If the calculated volumes are less than 50 μL or greater than 400 μL, consult the teaching faculty with how to move forward.
- Retrieve your CometChip from the 4 °C cooler.
- You will also need to gather one glass plate, one 96-well bottomless plate, and four 1.5" binder clips from the front bench.
- Remove your CometChip from the 1x PBS and place it, gelbond side down, on the glass plate.
- Press the 96-well bottomless plate onto the CometChip so that the wells line up with your labeling as shown in the diagram on the right.
- Be sure to press the top of the 96-well bottomless plate onto the CometChip. If you are unsure which side is the top, please ask the teaching faculty.
- Do not move the 96-well bottomless plate while it is on the CometChip as you will damage the agarose and the microwells.
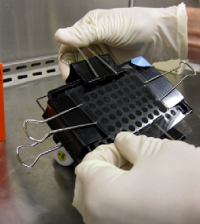
- Use the binder clips to secure the 96-well bottomless plate to the glass plate, thus creating a 'sandwich' with your CometChip in the center.
- Fasten the binder clips to the very edge of 96-well bottomless plate as shown in the image below.
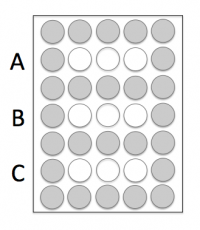
- You will load into the white wells and the grey wells should remain empty.
- Add 50 μL of 1x PBS to the Condition A wells.
- Add the appropriate volume of your cell suspension (calculated in Step #1) to the Condition B and C wells.
- Incubate your CometChip in the 37 °C incubator in the main laboratory for 15 min.
- After the incubation, complete a wash step to remove excess cells that are not within the microwells of your CometChip. Read all the bullets below before proceeding!
- Carefully remove the binder clips and the 96-well bottomless plate.
- Alert the teaching faculty at this point! The wash step can be very temperamental and it is best to see a demonstration!
- With the CometChip on the glass plate, 'waterfall' ~5 mL of 1x PBS over the wells, which are now imprinted onto the agarose.
- Hold the glass plate with the CometChip such that Condition C is at the bottom.
- To waterfall the 1x PBS, hold the glass plate at a 45° angle over the dish that you used to store your CometChip.
- Pipet up ~5 mL of 1x PBS.
- Press the pipet tip onto the glass plate above your CometChip.
- As you expel the 1x PBS, quickly move the pipet tip from left-to-right.
- The 1x PBS should pass over the top of the CometChip and fall into the dish.
- Use a P200 tip attached to the pasteur pipet to aspirate the excess liquid from your CometChip wells.
- Lightly touch the tip to the bottom of each imprinted well on the CometChip and immediately lift the tip from the agarose.
- Read through Steps #11-14 before continuing with the procedure.
- Retrieve one tube of molten 1% low melting point (LMP) agarose from the 42 °C waterbath.
- You will need to work quickly from this point as the LMP agarose will solidify as it cools.
- Using the P1000, pipet up 1000 μL of molten agarose from the tube.
- Hold the pipet tip over the top left well of your CometChip and as you expel the agarose move the pipet tip from left to right. Ensure that each row of your CometChip gets covered.
- The goal is to lightly cover the wells that contain cells, which will 'trap' the cells into the microwells.
- If the LMP agar 'fell' off the CometChip in any areas during this process, it is important to 'fill in' those portions of the CometChip. Please alert the teaching faculty if you experience any difficulties!
- Leave your CometChip undisturbed on the benchtop for 3 min then carefully move it to the 4 °C cooler for 5 min to ensure the LMP agarose solidifies.
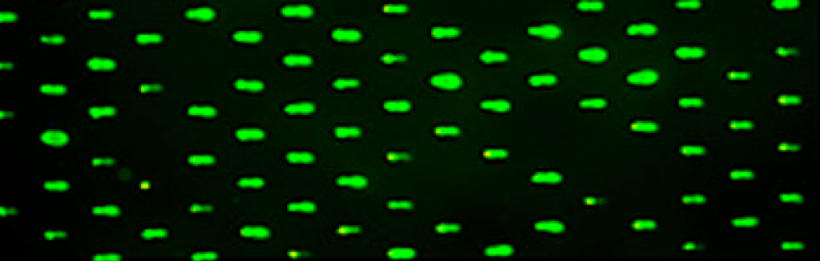
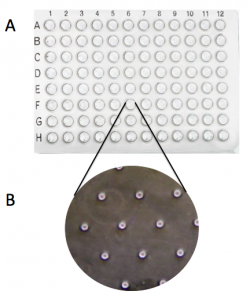
- Use the microscope in the main laboratory to image your CometChip. You will use these images to determine: 1. the number of microwells that are loaded / total number of microwells in the frame and 2. the number of cells / microwell.
- After imaging, obtain a fresh dish and a 30 mL aliquot of alkaline lysis buffer from the front laboratory bench.
- Carefully move your CometChip into the dish.
- Add the aliquot of alkaline lysis solution by slowly pouring it into a corner of the dish rather than directly over the CometChip.
- Be sure the CometChip is completely submerged and not floating in the dish.
- Label the dish with your team and section information.
- Move your dish into the 4 °C cooler for 60 min.
- Remove the alkaline lysis solution by pouring from the corner of the dish.
- Incubate the CometChip in 20mL of neutralization buffer for 15min. Repeat this step once more.
- Obtain a 30 mL aliquot of SYBR gold DNA stain from the front laboratory bench and add it to the dish containing your CometChip.
The teaching faculty will transfer your CometChip from the SYBR gold DNA stain solution into 1X PBS at ~24 hr.
Part 4: Practice statistics analysis
Review these data from an experiment where cells were exposed to increasing amounts of radiation. Your goal is to determine if a statistically significant amount of DNA damage was induced. For the purpose of this exercise, the values in the spreadsheet are in arbitrary units of 'DNA damage', where the higher numbers indicate more damage.
Recall the lecture delivered by Dr. Lyell on statistical analysis – how will you convince someone that the DNA damage was significant? You may find this spreadsheet, originally created by Prof. Bevin Engelward and modified by the 20.109 staff, helpful for this exercise. At a minimum, you should create a bar plot of the data with 95% confidence intervals and indicate if there is a statistically significant difference (i.e. provide a p-value) between conditions. Place this plot in your Benchling notebook.
Reagents list
CometChip:
- agar, normal melting point (Invitrogen)
- GelBond film (Lonza)
- 1 well dish (VWR)
- agar, low melting point (Invitrogen)
- bottomless 96-well plates (VWR)
- SYBR gold DNA stain, diluted 1:1000 in 1X PBS (Thermo Fisher)
- phosphate buffered saline (VWR)
- Alkaline Lysis Solution (2.5M NaCl, 100mM Na2EDTA, 10mM Tris, 1% Trition-X)
- Neutralization Buffer (0.4M Tris)
Cell culture:
- Mouse embryonic fibroblasts (MEF) cells (Engelward Laboratory)
- DMEM (Gibco)
- with 10% fetal bovine serum (Atlanta Biologicals)
- 100X antibiotic solution (Gibco)
- 10,000 U/mL Penicillin
- 10,000 U/mL Streptomycin
- Trypsin (Gibco)
- Trypan Blue
- Incubator maintains 37°C, 5% CO2 and 95% relative humidity
Next day: Evaluate cell loading results and prepare biochemical experiment