Difference between revisions of "20.109(S22):M2D4"
Noreen Lyell (Talk | contribs) (Created page with "<div style="padding: 10px; width: 820px; border: 5px solid #434a43;"> {{Template:20.109(S22)}} ==Introduction== ==Protocols== ==Reagents list== ==Navigation links==") |
Noreen Lyell (Talk | contribs) (→Protocols) |
||
| Line 7: | Line 7: | ||
==Protocols== | ==Protocols== | ||
| + | |||
| + | |||
| + | ====Confirm sgRNA_target with sequencing==== | ||
| + | |||
| + | The sgRNA_target sequence that was inserted into the expression plasmid was confirmed using DNA sequencing. The invention of automated sequencing machines has made sequence determination a relatively fast and inexpensive process. The method for sequencing DNA is not new but automation of the process is recent, developed in conjunction with the massive genome sequencing efforts of the 1990s and 2000s. At the heart of sequencing reactions is chemistry worked out by Fred Sanger in the 1970s which uses dideoxynucleotides, or chain-terminating bases. These chain-terminating bases can be added to a growing chain of DNA but cannot be further extended. Performing four reactions, each with a different chain-terminating base, generates fragments of different lengths ending at G, A, T, or C. The fragments, once separated by size, reflect the DNA sequence due to the presence of fluorescent dyes, one color linked to each dideoxy-base. The four colored fragments can be passed through capillaries to a computer that can read the output and trace the color intensities detected. | ||
| + | |||
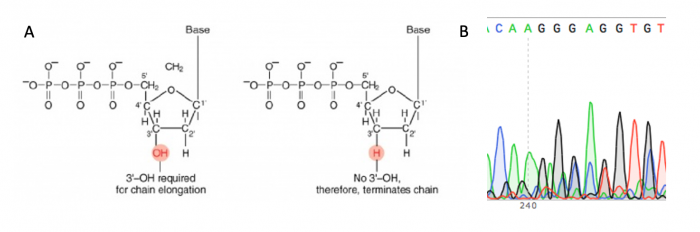
| + | [[Image:Fa20 M3D2 sanger sequencing.png|thumb|center|700px|'''Principles of Sanger sequencing.''' A. Chain-terminating bases are used to halt the DNA synthesis reaction at different lengths and attach a fluorophore that is used to determine the sequence of the DNA strand. B. The sequence of the DNA strand is determined using the fluorescent signature associated with each length of DNA in the reaction, this is visualized as a chromatogram.]] | ||
| + | |||
| + | <br style="clear:both;"/> | ||
==Reagents list== | ==Reagents list== | ||
==Navigation links== | ==Navigation links== | ||
Revision as of 14:16, 21 January 2022
Introduction
Protocols
Confirm sgRNA_target with sequencing
The sgRNA_target sequence that was inserted into the expression plasmid was confirmed using DNA sequencing. The invention of automated sequencing machines has made sequence determination a relatively fast and inexpensive process. The method for sequencing DNA is not new but automation of the process is recent, developed in conjunction with the massive genome sequencing efforts of the 1990s and 2000s. At the heart of sequencing reactions is chemistry worked out by Fred Sanger in the 1970s which uses dideoxynucleotides, or chain-terminating bases. These chain-terminating bases can be added to a growing chain of DNA but cannot be further extended. Performing four reactions, each with a different chain-terminating base, generates fragments of different lengths ending at G, A, T, or C. The fragments, once separated by size, reflect the DNA sequence due to the presence of fluorescent dyes, one color linked to each dideoxy-base. The four colored fragments can be passed through capillaries to a computer that can read the output and trace the color intensities detected.