Difference between revisions of "20.109(S22):M1D2"
Noreen Lyell (Talk | contribs) (→Part 1: Analyze SMM results) |
Noreen Lyell (Talk | contribs) (→Part 1: Analyze SMM results) |
||
| Line 38: | Line 38: | ||
'''Quantify fluorescence on 635 nm channel''' | '''Quantify fluorescence on 635 nm channel''' | ||
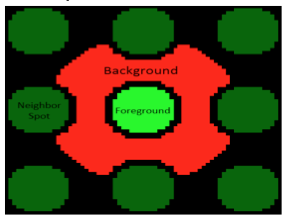
| − | [[Image:Sp22 M1D2 P1 quantify 635nm.png|thumb|right|300px]] After the grid pattern is aligned, the fluorescence is measured on the 635 nm channel. In this, the slide is 'read' pixel by pixel and a numerical value is assigned that indicates the intensity of the signal measured at a particular pixel as illustrated in the image to the right. The computer software uses the numerical values to calculate the signal-to-noise ratio. | + | [[Image:Sp22 M1D2 P1 quantify 635nm.png|thumb|right|300px]] After the grid pattern is aligned, the fluorescence is measured on the 635 nm channel. In this, the slide is 'read' pixel by pixel and a numerical value is assigned that indicates the intensity of the signal measured at a particular pixel as illustrated in the image to the right. The computer software uses the numerical values to calculate the signal-to-noise ratio (<math> SNR </math>). |
| + | |||
| + | <center> | ||
| + | <math> SNR = \frac{\mu _{foreground}-\mu _{background}}{\theta_{background} | ||
| + | </center> | ||
<font color = #4a9152 >'''In your laboratory notebook,'''</font color> complete the following: | <font color = #4a9152 >'''In your laboratory notebook,'''</font color> complete the following: | ||
Revision as of 18:11, 18 January 2022
Introduction
Though you may be able to qualitatively visualize the spots that appear to emit more fluorescence, it is important to complete quantitative analysis that supports your observations. The microarrayer reads the fluorescence signals emitted from the surface of the SMM slide at two excitation wavelengths. As noted previously, the 532 nm wavelength was used to excite fluorescein, which was printed in an 'X' pattern to assist with alignment. The 635 nm wavelength was used to excite Alexa Fluor 647; which would be associated with TDP43-RRM12 bound to a small molecule on the slide. A hit denotes a spot on the slide that emits a red fluorescence signal significantly higher than the background fluorescence level. In terms of protein binding, a hit denotes that the TDP43-RRM12 protein is bound to a small molecule and is therefore localized to a specific position on the slide. You will analyze the fluorescence signal collected by the microarray scanner using a value termed the robust z-score.
The robust z-score differentiates signal from noise by providing a value that represents the intensity of a signal above background. In the case of the SMM experiment, the intensity of a fluorescent signal above the background fluorescence is calculated. To do this the fluorescence emitted across the entire slide is grouped to define the Median Absolute Deviation (MAD), which is is a measure of the variability of a univariate dataset. Though beyond the scope of this class, the equation for calculating the robust z-score assigns a value for how much more intense the fluroescent signal at a spot is over background. The higher the value, the more different the signal from background.When the SMM slides were imaged, the microarrayer also produced a GAL file, or GenePix Array List. The GAL file contains information about where each spot was printed, and what compound was printed there. However, the relationship between the GAL file and the actual contact of the print head is very imprecise. Instead, the fluorescein guide spots are used to align the array in the GAL file to the true print location for each pin. Following the alignment, the software quantifies the fluorescence at 635 nm within the deposition region of each spot (foreground) and the fluorescence immediately outside of this region, where nothing was printed (background) as illustrated in the image to the right. These values are used to calculate the robust z-score. From the robust z-score, you can determine the associated probability that the observed fluorescence occurred by chance, and if this probability is sufficiently low, we call the small molecule a 'hit'.
After hits are identified via the robust z-score, the data are examined by-eye using the criteria discussed in the prelab. In the exercises below you will review this process to gain a better understanding for how hits the hits identified in the SMM screen that targeted TDP43-RRM12 were validated.
Protocols
Part 1: Analyze SMM results
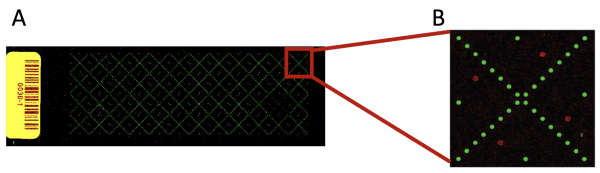
In the previous laboratory session, you reviewed how the SMM screen for TDP43-RRM12 was completed and how the slides were imaged for analysis. Prior to performing the SMM experiment, small molecule compounds were printed onto slides using the method described in the M1D1 Introduction. Each slide was printed with ~12,000 spots! In addition to ~4,200 small molecules (printed in duplicate), fluorescein and DMSO spots were included on every slide. The fluorescein spots, or sentinel spots, are used to align a grid pattern to the slide so the small molecule compounds at each spot can be identified. The DMSO spots are a negative control. Each slide is arranged as depicted below. The fluorescein spots are printed in an "X" pattern across the slide (panel A). Each "X" section represents a block within the slide (panel B).
The goal for today is to more thoroughly explore the analysis steps that were used to identify the hits for TDP43-RRM12. In this exercise you will consider the importance of each of the four steps listed below in the identification of hits.
- Align printed small molecule spots using fluorescence on 532 nm channel.
- Quantify fluorescence on 635 nm channel.
- Identify hits with improbably high fluorescence.
- Complete 'by-eye' analysis of hits.
Align printed small molecule spots using fluorescence on 532 channel
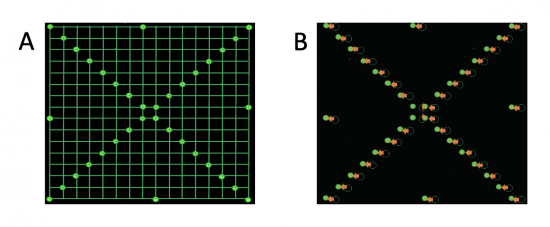
To process the slide images, a grid pattern (panel A) is aligned to each block using the fluorescence measured on the 532 channel. In the example to the right (panel B), the closed green spots represent the fluorescence signal on the 532 channel and the open green circles represent the software alignment. As you can see, the software misaligned the "X" pattern for this block. The orange arrows indicate the manual manipulation completed by the user to correct the alignment.In your laboratory notebook, complete the following:
- What is the source of the fluorescence measured on the 532 nm channel?
- Why is it important that the "X" pattern is correctly aligned for each block?
Quantify fluorescence on 635 nm channel
After the grid pattern is aligned, the fluorescence is measured on the 635 nm channel. In this, the slide is 'read' pixel by pixel and a numerical value is assigned that indicates the intensity of the signal measured at a particular pixel as illustrated in the image to the right. The computer software uses the numerical values to calculate the signal-to-noise ratio ($ SNR $).