20.109(F22):M2D2
Introduction
Today you will isolate the expressed PfFKBP35 protein from the bacterial cells. Remember that the PfFKBP35 gene sequence was synthesized such that a 6xHis-tag was added to the DNA sequence. The resultant protein is therefore His-tagged. Histidine has several interesting properties, notably its near-neutral pKa, and His-rich peptides are promiscuous binders, particularly to metals. (For example, histidine side chains help coordinate iron molecules in hemoglobin.)
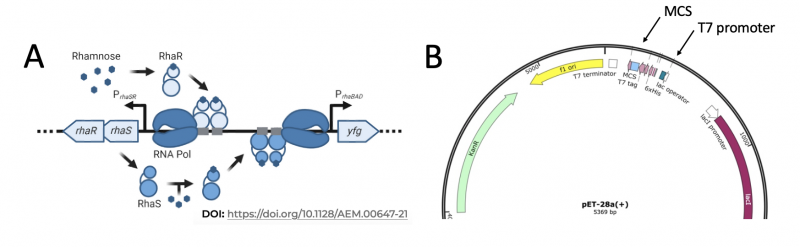
To induce production of PfFKBP35 protein from the expression plasmid that was 'cloned' in the previous laboratory session, rhamnose was used to induce expression in KRX E. coli bacterial cells. The use of rhamnose to induce protein expression is based on the native rha operon used for rhamnose metabolism in bacterial cells. The native rha operon is involved in rhamnose metabolism in bacterial cells and composed of five genes: rhaR, rhaS, rhaB, rhaA, and rhaD. RhaR and RhaS have a regulatory role as activators of RhaBAD, which are directly responsible for the metabolism of rhamnose. When rhamnose is present, a regulatory cascade is triggered in which RhaR is activated. Activated RhaR associates with RNA polymerase to drive transcription of more rhaR and rhaS (see Panel A in image below). RhaS then binds rhamnose and associates with RNA polymerase to drive transcription of rhaB, rhaA, and rhaD from the rhaBAD promoter (rhaPBAD). In the KRX cells used to produce PfFKBP35, the rhaBAD genes are absent and the gene that encodes T7 RNA polymerase is controlled by rhaPBAD. This is represented by ‘yfg’ or ‘your favorite gene’ in Panel A of the image below.
So, why do we want to control expression of T7 RNA polymerase? As you learned in the previous laboratory session, the PfFKBP35 gene was cloned into the pET-28a expression vector. In this vector, a T7 promoter (PT7) is located upstream of where the PfFKBP35 gene was cloned and is used to drive expression (see Panel B in image below). Therefore, the T7 RNA polymerase that is encoded in the KRX genome and controlled by rhamnose via RhaSR transcribes PfFKBP35 from the PT7 in the pET-28a_PfFKBP35 expression plasmid!
By engineering the native rha operon, researchers were able to develop a powerful tool that enables the control of gene expression using an inducer molecule. Because RhaSR are only active when rhamnose is present, rhamnose can be used as an inducer to control the production of a protein of interest.
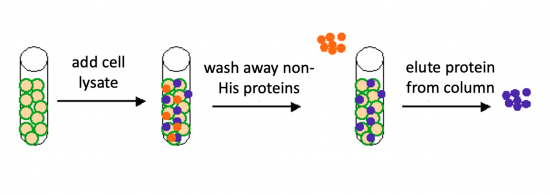
To purify the PfFKBP35 proteins present in the bacterial cell, you will use a nickel-agarose resin. The 6xHis-tag on the PfFKBP35 protein will bind to the nickel-coated resin, while the other cellular protein will pass through the resin. Remember, the KRX cells are not only producing the PfFKBP35 protein, but also the proteins needed for cellular function and survival. Imidazole is a compound that is also able to bind to nickel and washing the resin with a low concentration solution promotes competition for binding between the imidazole and bound proteins for the nickel-coated resin. Proteins that are non-specifically bound will have a lower affinity for the nickel than imidazole and be washed from the column, whereas the 6xHis-tagged PfFKBP35 will remain adhered to the nickel-agarose resin. To elute the PfFKBP35 protein from the nickel-coated resin, a high concentration of imidazole is used to out-compete the 6xHis-tag for binding.
Protocols
Part 1: Induce expression of PfFKBP35
For timing reasons, the induction steps were completed prior to class. So you understand how the cell pellets you will use for the protein purification, the steps are shown in the video and protocol below.
To ensure the steps included below are clear, please watch the video tutorial linked here: [Bacterial Induction]. The steps are detailed below so you can follow along!
- Inoculated 5 mL of LB media containing 50 μg/mL kanamycin with a colony of KRX cells transformed with pET-28a_FKBP35.
- Incubated the culture overnight at 37 °C with shaking at 220 rpm.
- Dilute the overnight culture 1:100 in 50 mL of fresh LB media containing 50 μg/mL kanamycin.
- Incubate at 37 °C until the OD600 = 0.6 with shaking at 220 rpm.
- To induce PfFKBP35 protein expression, add rhamnose to a final concentration of 0.1%.
- Incubate overnight at room temperature with shaking at 150 rpm.
- To harvest the cells, centrifuge the culture at 4000 g for 15 min at 4 °C.
- Cell pellets were flash frozen in liquid nitrogen, then stored at -80 °C until used for purification.
In your laboratory notebook, complete the following:
- Calculate the volume of kanamycin stock that was added to the LB broth in Step #1. In Step #3.
- Concentration of kanamycin stock = 50 mg/mL.
- Calculate the volume of rhamnose stock that was added to the LB broth in Step #5.
- Concentration of rhamnose stock = 20% (w/v).
Part 2: Purify PfFKBP35 protein
To ensure the steps required for purifying the protein are clear, the Instructor will provide a live demonstration of this process.
Lyse KRX cells expressing pET-28a_FKBP35
- Retrieve a conical tube containing a KRX pET-28a_FKBP35 cell pellet from the -80 °C freezer and leave it on your bench to thaw.
- To lyse the cells, resuspend the pellet in B-Per bacterial extraction reagent at 2 mL / cell pellet.
- To the cell suspension add:
- lysozyme at 2 μL / mL of B-Per bacterial extraction reagent
- DNase at 2.5 μL / mL of B-Per bacterial extraction reagent
- 1 protease inhibitor tablet
- Solubilize the cell pellet in lysis buffer and vortex to mix.
- Incubate cell pellet in lysis buffer at room temperature for 15 min on nutator.
- To pellet the cell debris, centrifuge the lysate at 3,000 rcf for 45 min at 4 °C.
- Complete Part 3: Electrophorese confirmation digests during the centrifugation.
Prepare Ni-NTA affinity column
- Obtain a 500 μL aliquot of 50% slurry (Ni-NTA resin) and mix the slurry by inverting the tube several times.
- The slurry is the Ni-NTA column matrix.
- Centrifuge the slurry for 30 sec then remove the supernatent.
- To wash the slurry, add 500 μL of 1X PBS and invert the tube 3 times.
- Add the slurry to the column and allow the 1X PBS to run through the column.
- Be sure a beaker is placed under the column to collect the waste!
- When the PBS has flowed through, cap the bottom and the top of the column until you are ready to add the cell lysate.
Purify PfFKBP35 from cell lysate
- Transfer the supernatent from the centrifuged cell lysate to a fresh conical tube.
- Label the microcentrifuge tube containing the cell pellet as "pellet" and give it to the Instructor! This pellet will be used later when protein expression and purity are examined.
- Aliquot 30 μL of the supernatent (from Step #1) to a fresh microcentrifuge tube.
- Label the microcentrifuge tube containing the aliquot as "lysate" and give it to the Instructor! This aliquot will be used later when protein expression and purity are examined.
- Pipet the supernatent into the prepared Ni-NTA affinity column.
- Be sure that the bottom of the column is capped!
- Pipette the resin / cell supernatent to mix and incubate at room temperature on a nutator for 15 min.
- Hold a microcentrifuge tube under the column, then remove the bottom cap from the column and collect the liquid that leaves the column.
- Label the microcentrifuge tube as "flowthrough" and give it to the Instructor! This aliquot will be used later when protein expression and purity are examined.
- To wash the Ni-NTA affinity column, add 10 mL of wash buffer.
- Hold a microcentrifuge tube under the column, then remove the bottom cap from the column and collect ~250 μL of the liquid that leaves the column.
- Label the microcentrifuge tube as "wash" and give it to the Instructor! This aliquot will be used later when protein expression and purity are examined.
- To elute the PfFKBP35 protein from the affinity column, add 1 mL of elution buffer and incubate at room temperature for 5 min.
- Hold a microcentrifuge tube under the column, then remove the bottom cap from the column and collect the entire 1 mL of the liquid that leaves the column.
- Label the microcentrifuge tube as "elution" and give it to the Instructor! This aliquot will be used later when protein expression and purity are examined.
- Lastly, resuspend the slurry from the Ni-NTA affinity column in 250 μL 1X PBS and transfer to a fresh microcentrifuge tube.
- Label the microcentrifuge tube as "slurry" and give it to the Instructor! This aliquot will be used later when protein expression and purity are examined.
In your laboratory notebook, complete the following:
- At several steps in the protein purification procedure, samples are collected that will be used later when protein expression and purity are examined. Consider why each of the samples listed below are saved as controls to measure the success of the purification.
- The pellet from Step #1.
- The lysate from Step #2.
- The flowthrough from Step #6.
- The wash from Step #7.
- The slurry from Step #9.
- What is occurring during the incubation in Step #4?
Part 3: Electrophorese confirmation digests
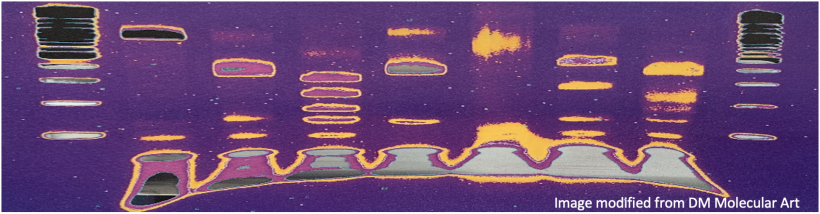
Electrophoresis is a technique that separates large molecules by size using an applied electrical field and a sieving matrix. DNA, RNA and proteins are the molecules most often studied with this technique; agarose and acrylamide gels are the two most common sieves. The molecules to be separated enter the matrix through a well at one end and are pulled through the matrix when a current is applied across it. The larger molecules get entwined in the matrix and are stalled; the smaller molecules wind through the matrix more easily and travel farther away from the well. The distance a DNA fragment travels is inversely proportional to the log of its length. Over time fragments of similar length accumulate into “bands” in the gel. Higher concentrations of agarose can be used to resolve smaller DNA fragments.
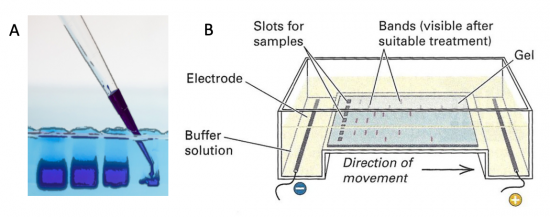
DNA and RNA are negatively charged molecules due to their phosphate backbone, and they naturally travel toward the positive electrode at the far end of the gel. Today you will separate DNA fragments using an agarose matrix. Agarose is a polymer that comes from seaweed. To prepare these gels, agarose and 1X TAE buffer (Tris base, acetic acid, and EDTA) are microwaved until the agarose is melted and fully dissolved. The molten agar is then poured into a horizontal casting tray, and a comb is added. Once the agar has solidified, the comb is removed, leaving wells into which the DNA samples can be loaded.
For the digests that were prepared in the previous laboratory session, a 1% agarose gel with SYBR Safe DNA stain was used to separate the DNA fragments in the four digest reactions. In addition, a well was loaded with a molecular weight marker (also called a DNA ladder) to determine the size of the fragments.
To ensure the steps included below are clear, please watch the video tutorial linked here: [DNA gel electrophoresis].
- Add 5 μL of 6x loading dye to the digests.
- Loading dye contains bromophenol blue as a tracking dye, which enables you to follow the progress of the electrophoresis.
- Glycerol is also included to weight the samples such that the liquid sinks into well.
- Flick the eppendorf tubes to mix the contents, then quick spin them in the microfuge to bring the contents of the tubes to the bottom.
- Load 25 μL of each digest into the gel, as well as 5 μL of 1kb DNA ladder.
- Be sure to record the order in which you load your samples!
- To load your samples, draw the volume listed above into the tip of your P200 or P20. Lower the tip below the surface of the buffer and directly over the well. Avoid lowering the tip too far into the well itself so as to not puncture the well. Expel your sample slowly into the well. Do not release the pipet plunger until after you have removed the tip from the gel box (or you'll draw your sample back into the tip!).
- Once all the samples have been loaded, attach the gel box to the power supply and electrophorese the gel at 125 V for 45 minutes.
- Lastly, visualize the DNA fragments in the agarose gel using the gel documentation system.
Reagents list
- Luria-Bertani broth (LB) (from Difco)
- kanamycin (from Sigma)
- rhamnose (from Sigma)
- 2x B-Per bacterial protein extraction reagent (from ThermoFisher)
- lysozyme (from Sigma)
- DNase (from Sigma)
- protease inhibitor tablet (from Sigma)
- phosphate saline buffer (PBS) (from VWR)
- Ni-NTA agarose (from Qiagen)
- Wash buffer: 100 mM HEPES (pH = 7.4), 500 mM NaCl, 50 mM imidazole
- Elution buffer: 100 mM HEPES (pH = 7.4), 500 mM NaCl, 250 mM imidazole
- imidazole (from Sigma)
Next day: Assess purity and concentration of purified protein