20.109(F21):Module 2
Contents
Module 2: Drug discovery
Malaria is a mosquito-borne disease caused by Plasmodium parasites. This life-threatening disease is estimated to be responsible for over 409,000 deaths from 229 million cases worldwide in 2019. The impact of malaria is disproportionately seen in Africa, where 95% of malaria cases and 95% deaths occurred (World Health Organization, World Malaria Report, 2020).
In this module, you will explore drug discovery by focusing on a specific infectious disease, malaria, as an example of how small molecules can be used in therapeutics. The process by which new therapeutics are discovered, optimized and tested to ensure safety and effectiveness is lengthy. Here, we will focus on the discovery phase and thinking through what makes a good therapeutic. This is ideally grounded by molecular level understanding of the disease process, and knowledge of specific targets or pathways that can be manipulated to improve health outcomes. In some instances, however, effective small molecule therapeutics can be developed despite incomplete biological understanding.
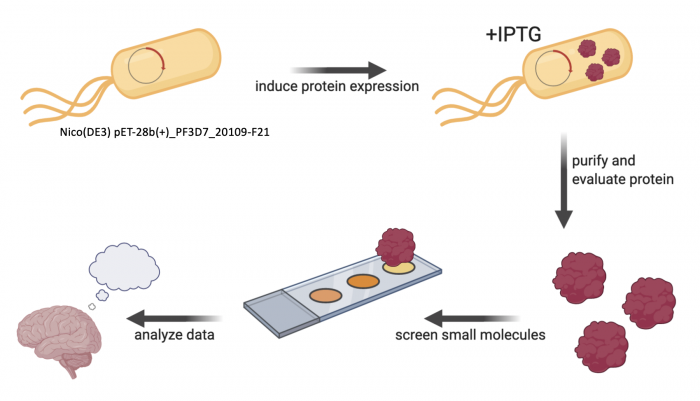
Small molecule screens are foundational to drug discovery. In this module, you will learn about small-molecule microarray (SMM) technology and how this can be used to identify molecules that putatively bind to a protein from Plasmodium falciparum that is essential for its survival. The Niles Laboratory has been working to identify and validate essential proteins of unknown function(s) from P. falciparum as potential targets for drug discovery. Through these efforts, multiple proteins have been selected as candidate targets, and are being recombinantly expressed to use in SMM screens to identify small molecules binding hits.
The protein you will work with was identified via this process. Though much remains to be learned about its function(s) in P. falciparum, available biological data suggest that it could be a valid therapeutic target. SMM and other screens have identified several interesting binding hits, but these need to be confirmed in secondary binding assays before selecting candidates to prioritize for additional characterization and optimization. As part of this module, you will have the opportunity to perform quantitative binding assays, as well as analyze and interpret these data to help inform which molecules may be stronger candidates for further study.
Research goal: Identify small molecules that bind to the PF3D7_20109-F21 protein in Plasmodium falciparum using small-molecule microarray
Lab links: day by day
M2D1: Review small molecule microarray (SMM) experiment and results
M2D2: Complete in silico cloning of protein expression plasmid
M2D3: Perform protein purification protocol
M2D4: Assess purity and concentration of purified protein
M2D5: Perform secondary assay to test putative small molecule binders
M2D6: Complete data analysis for secondary assay
M2D7: Plot and interpret data from secondary assay
Major assignments
Journal club presentation
Research article
References
A method for the covalent capture and screening of diverse small molecules in a microarray format. Nature Protocols. (2006) 1:2344-2352.
Recent discoveries and applications involving small-molecule microarrays. Chemical Biology. (2014) 18:21-28.
The Strep-tag system for one-step purification and high-affinity detection or capturing of proteins. Nature Protocols. (2007) Vol.2. No.6:1528-1535.
Uncovering the essential genes of the human malaria parasite Plasmodium falciparum by saturation mutagenesis. Science. (2018) 360:506.
- Reference for mutagenesis index score: plot linked here
- Plot shows ranked order of all Plasmodium falciparum genes from most essential (left) to dispensable (right) along the x-axis. A piggyBAC insertion mutagenesis screen (mobile element that inserts into genes to disrupt production of their encoded protein) was used to generate the plotted data.