20.109(F17):Evaluate cell loading results (Day3)
Contents
Introduction
Today you will continue to examine the data collected for your cell loading experiment, and then use these results to probe the effects of biochemical factors on genomic stability. DNA damage is defined as any change in the chemical structure of the molecule, including breaks in the backbone, missing basepairs, and altered basepairs. Damaged DNA does not mean mutated DNA! Though both instances relay that the DNA has been changed, a mutation is defined as a change in the sequence.
In lecture, you reviewed exogenous factors that lead to DNA damage (i.e. ultraviolet rays, smoking, etc.). Naturally occurring DNA damage results from native cell processes, such as metabolism. In this module, we will examine the effects of oxidative and alkylating agents on genomic stability using the CometChip assay.
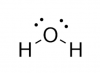
Oxidative agent: hydrogen peroxide (H2O2)Normal cell tissues have a basal level of DNA damage due to cell processes involved in cellular metabolism. For example, electrons can escape the electron transport chain and result in the formation of superoxide. Furthermore, defense mechanisms employed to protect the host from bacterial infection involved the release of reactive oxygen species. These reactive oxygen species are implicated in causing more than 20 types of DNA base lesions.
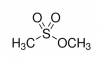
Alkylating agent: methyl methanesulfonate (MMS)Exposure to alkylating agents occurs via the environment, food, and even cancer chemotherapeutic treatments. Alkylation is the transfer of an alkyl group from one molecule to another. In the context of DNA damage, most alkylating agents methylate DNA forming adducts at the N- and O- atoms in the bases. This change in the DNA results in double-strand breaks.
Protocols
Part 1: BE Communication Lab workshop
Our communication instructors, Dr. Sean Clarke and Dr. Prerna Bhargava, will join us today for a workshop on designing effective figures and captions.
Part 2: Image cell loading experiment
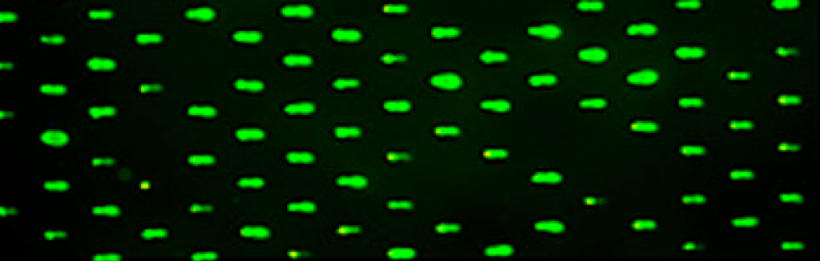
In the previous laboratory session you used the light microscope in the teaching laboratory to image your CometChip and determine the number of wells that were loaded and potentially how many cells were in each of the wells. Today you will use a fluorescent microscope in the Engelward Laboratory to confirm that enough cells were loaded to allow for the visualization of the DNA in the microcells using a florescent DNA dye. This will also allow you to see the microscope in the Engelward Laboratory, which will be used for all CometChip imaging in this module.
The teaching faculty will take one team at a time to the microscope in the Engelward Laboratory and image your CometChip. With the light microscope images, you determined the number of microwells that contained cells and potentially how many cells were present in the microwells. The images you examine today will give information on the number of wells that contain enough DNA to provide a fluorescent signal after staining. You can compare the number of microwells that contain cells from the light microscopy images to the number of wells that produce signal from the fluorescence microscopy images. Ideally, these numbers will be similar.
Part 3: Prepare CometChips to test biochemical factors
- Obtain a sheet of gelbond film from the laboratory bench at the front of the room. The paper is protecting the hydrophilic side of the gelbond film.
- Be sure to keep the paper associated with the gelbond film so you know which side is which.
- Use a Secureline II pen and a ruler to draw a 5.5 cm x 5.5 cm square on the hydrophobic side of the gelbond film. This square will encompass a 6 well x 6 well grid on a 96 well plate.
- Note: you are writing on what will be the bottom of the CometChip and may want to write backwards so the labels are clear when you look at the top of your CometChip.
- Prepare 20 mL of 1% normal melting point (NMP) agarose. Be careful as the agarose solution will be very hot!
- Calculate the amount of NMP agarose powder needed for a 1% w/v solution. Check your math with the teaching faculty before you continue.
- Obtain a small milk bottle from the front bench.
- Weigh out the appropriate amount of NMP agarose and add it to the milk bottle.
- Use a cylinder to measure 20 mL of 1x PBS and add it to the milk bottle with the NMP agarose powder.
- Swirl to mix.
- Rinse the cylinder with deionized water and place on rack above sink to dry.
- To melt the NMP agarose, microwave the solution for 20 seconds, swirl, then microwave for 3-second intervals until all crystals are in solutions. After each interval, remove the milk bottle and gently swirl while checking for unmelted agarose crystals. It is important that the solution does NOT boil as you will lose water to evaporation and the density of the agarose will be altered. If your solution starts to boil, immediately remove it from the microwave and gently swirl.
- When no more crystals are visible in the solution take the milk bottle to your bench.
- Obtain a small rectangle dish labeled "scraped lid" and the CometChip 'stamp' from the front bench.
- Add 2.5 mL of the agarose solution to the small dish, then quickly place the gelbond film in the dish with the marked hydrophobic side down.
- Add 13 mL of the agarose solution on top of the gelbond film.
- Slowly place the CometChip stamp on top of the agarose.
- Lower the bottom left of the stamp first, then slowly allow the stamp to 'roll' into the agarose. Be sure to leave the top right corner of the small dish accessible.
- Be careful not to introduce bubbles into the agarose and work quickly as the agarose will solidify as it cools.
- Allow the agarose to solidify, undisturbed, on your bench for 30 min.
- Meanwhile, rinse out your milk bottle in the sink with deionized water and hang on the dish rack to dry
- Add ~5 mL of 1x PBS to the small dish that contains your agarose CometChip.
- Pipet in the 1x PBS using the accessible corner.
- Slowly pull from one corner of the stamp to lift it away from your CometChip in the dish.
- If the CometChip sticks to the stamp, carefully peel it off using tweezers.
- Discard the PBS in the sink.
- Remove excess agarose from the perimeter of your CometChip using a razor blade.
- Clean the agarose from the bottom of your CometChip (gelbond side) using a Kimwipe.
- Place your CometChip in the small dish containing 1x PBS for storage at 4 °C until next time.
- Be sure the chip is completely submerged.
- Clean out your "scraped lid" to use as the lid of your dish.
- With the remaining time, begin to prepare a second CometChip. Each team will need a total of two CometChips for the biochemical testing experiment. If you do not have time to completely finish the second chip, the teaching faculty will finish the preparation procedure.
Part 4: Determine cell loading number for subsequent experiments
In a group discussion with the teaching faculty, you will assess the results of the class data from the CometChip loading experiments. The goal here is to determine which cell number to use when preparing your CometChip for the next experiments. Use the data you collected concerning the following: 1. number of wells loaded, 2. an estimate of number of cells per well, and 3. strength of DNA signal.
Be sure to include notes on the discussion and the values for cell loading number that you will use in your notebook!
Reagents list
CometChip:
- agar, normal melting point (Invitrogen)
- phosphate buffered saline
- GelBond film (Lonza)
- 1 well dish (VWR)
Next day: Test role of biochemical factors in genomic stability