20.109(S17):Purification of induced protein (Day2)
Contents
Introduction
Last time you used the lactose-analogue IPTG to induce expression of inverse pericam in BL21(DE3)pLysS bacteria. Today you will isolate your mutated IPC and the wild-type IPC from the bacterial cells. You will also begin characterizing your wild-type and mutant proteins.
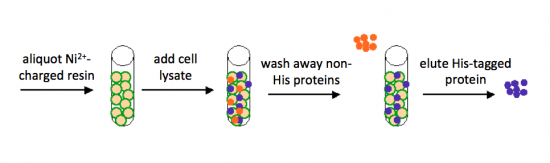
The bacterial expression vector we are using (pRSET) contains six histidine codons downstream of a bacterial promoter and in-frame with a start codon. Our resultant protein is therefore marked by the presence of these additional encoded residues, or His-tagged. Histidine has several interesting properties, notably its near-neutral pKa, and His-rich peptides are promiscuous binders, particularly to metals. (For example, histidine side chains help coordinate iron molecules in hemoglobin.)
Today we will use a nickel-agarose resin to separate our protein of interest (IPC) from the other proteins present in the bacteria. The His-tagged protein will preferentially bind to the nickel-coated beads, while proteins irrelevant to our purposes in Module 1 can be washed away. Remember, the BL21(DE3)pLysS cells are not only producing our protein, but also the proteins needed for cellular function and survival. Finally, a high concentration of imidazole (which is the side chain of histidine) can be used to elute the His-tagged inverse pericam by competition. Due to the inherent fragility of IPC, we will add several components to our protein extraction and purification reagents: bovine serum albumin (BSA), which is a protein stabilizer, and a cocktail of protease inhibitors.
Protocols
Part 1: Lyse BL21(DE3)pLysS pRSETb_FKBP12 cells
- Retrieve your BL21(DE3)pLysS pRSETb_FKBP12 cell pellet from the front laboratory bench and leave it on your bench to thaw.
- You will also obtain a cell pellet of uninduced cells that was prepared by the teaching faculty as a control to examine the IPTG induction step.
- Prepare 3 mL of lysis buffer.
- Use the information in the table below to calculate the volume of each stock reagent that is needed to prepare the lysis buffer with the specified final concentrations.
- Confirm your calculations with the teaching faculty, then prepare your lysis buffer solution.
- Weigh each cell pellet using the balance and add 1 mL of lysis buffer per g of cell pellet.
- Note: the weight of the 50 mL conical tube is XX g.
- Resuspend each pellet completely in the lysis buffer then transfer the cell suspension to a 2 mL eppendorf tube.
- Add lysozyme (stock concentration of 50 mg/mL) to each cell suspension such that the final concentration is 300 μg/mL.
- Incubate in the 4 °C cooler for 1 hr on the nutator.
| Stock reagent | Final concentration of stock reagent in lysis buffer | Volume of stock reagant |
|---|---|---|
| 1 M Tris (pH = 7) | 50 mM | |
| 1 M NaCl | 150 mM | |
| 40% glycerol | 10% | |
| 1 M DTT | 1 mM | |
| 1 M AEBSF | 1 mM | |
| H2O | add for a total of 3 mL of lysis buffer |
Part 2: Prepare Ni-NTA affinity column
Keep all buffers on ice when not in use. All spins should be performed at 1000 rcf (3300 rpm) for 1 minute.
- The following buffers are aliquoted and located at the front laboratory bench:
- Ni-NTA His-bind resin
- 1X Ni-NTA Bind Buffer (50 mM NaH2PO4, pH 8.0; 300 mM NaCl; 10 mM imidazole)
- 1X Ni-NTA Wash Buffer (50 mM NaH2PO4, pH 8.0; 300 mM NaCl; 20 mM imidazole)
- 1X Ni-NTA Elute Buffer (50 mM NaH2PO4, pH 8.0; 300 mM NaCl; 250 mM imidazole)
- Note: Two special waste streams should be created for this affinity purification procedure, (1) nickel waste for the 50% slurry, and (2) imidazole waste for the Bind, Wash, and Elute Buffers.
- Gently mix the Ni-NTA His-bind resin to fully resuspend it, then distribute 400 μL of the resin to each of two 2 mL centrifuge tubes.
- Label one tube as wild type and the other as mutant.
- Add 1.6 mL (2 x 800 μL) of 1X Ni-NTA Bind Buffer to the Ni-NTA His-bind resin.
- Resuspend the resin by pippeting the solution up and down several times (10-15), then centrifuge (see conditions above).
- Carefully remove the supernatant and discard it in the appropriate waste stream.
- Equilibrate column with PBS for 30 min at 4 C???
Part 3: Purify FKBP12 protein
- Retrieve your lysed cell pellets from the 4 °C cooler.
- Transfer 15 μL from each tube into labeled 1.5 mL eppendorf tubes and give the aliquots to the teaching faculty.
- You will use these aliquots during the next laboratory session to examine protein yield using polyacrylamide gel electrophoresis (PAGE).
- Add XX μL MgCl2 and 10 μL of DNase to the tube that contains the IPTG-induced sample.
- You will only complete the purification protocol for the IPTG-induced sample!
- Incubate for 30 min in the 4 °C cooler.
- Centrifuge your sample for 30 min at 16,000 rcf in the 4 °C cold room.
- Alert the teaching faculty when you are ready to centrifuge your sample and you will be escorted to the cold room.
- Check that the supernatent in your sample is clear with little to no 'cloudiness'.
- Load the supernatent onto the column your prepared in Part 2.
- Obtain an aliquot of PBS buffer containing 10 mM imidazole from the front laboratory bench.
- Wash the column by adding 1.5 mL of the PBS buffer containing 10 mM imidazole to the column and invert, then centrifuge at 3.3 rpm for 1 min.
- Transfer the flow-through to a fresh, well-labeled 2 mL eppendorf tube.
- Repeat Step #9 a total of three times.
- Elute your FKBP12 protein by adding 500 μL of PBS buffer containing 250 mM imidazole to the column and invert, then centrifuge at 3.3 rpm for 1 min.
- Transfer the purified protein sample (the flow-through) to a fresh, well-labeled 1.5 mL eppendorf tube.
- ADD procedure for dialysis setup
- Give your flow-through samples from Step #9 and your used purification column to the teaching faculty.
Your purified protein solution will dialyze for ~18 hr in PBS buffer to remove the imidazole as it may interfere with the small molecule screen. Following dialysis, the teaching faculty will transfer your purified protein solution to a fresh eppendorf tube and store it at XX °C until your next laboratory session.
Reagents
Next day: Evaluation of purified protein
Previous day: In silico cloning and induction of protein expression