Difference between revisions of "20.109(S14):Microbial DNA extraction (Day1)"
(→Introduction) |
MAXINE JONAS (Talk | contribs) m (28 revisions: Transfer 20.109(S14) to HostGator) |
||
| (23 intermediate revisions by 4 users not shown) | |||
| Line 3: | Line 3: | ||
<div style="padding: 10px; width: 640px; border: 5px solid #FF6600;"> | <div style="padding: 10px; width: 640px; border: 5px solid #FF6600;"> | ||
| − | ==Introduction== | + | ==Introduction== |
| − | + | Please begin by reading the [[20.109%28S14%29:Module_1 | '''Module 1 overview'''.]] | |
| − | + | Today we'll begin our primary experiment, a phylogenetic analysis of bird gut microbiota. Through birds' stool, their resident microbes can be transferred to the environment and in some cases infect other animals. Perhaps the most well-known pathogenic avian microbe with zoonotic potential (potential for inter-species transmission) is the flu virus. For your safety, all the samples we will work with have been screened to exclude those carrying human���pathogenic flu strains. However, we will be able to mine much of the same intellectual content that we could were we studying flu directly. | |
| − | + | Investigations in disease ecology, or "the ecological study of host-pathogen interactions within the context of their environment and evolution" ([http://www.nature.com/scitable/knowledge/library/disease-ecology-15947677 '''via Scitable''']), help determine how pathogens transmit and cause disease, persist, and evolve in host organisms as different as humans and birds. Pathogen, host, and environment all play roles in defining the natural history of disease. In studies of pathogens that cause zoonotic disease, researchers are particularly interested in defining major influences on pathogen distribution, transmission, and evolution. The Runstadler lab currently studies the disease ecology of influenza viruses in several groups of birds and other animals (see for example [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3115932/ '''this paper'''] and [http://www.ncbi.nlm.nih.gov/pubmed/22192630 '''this one''']), a research area that can provide information useful for predicting the next flu pandemic and designing effective vaccines that will be produced in time. Researchers track viral mutations/evolution, infection of different bird species (including co-infection by multiple strains), and the trafficking patterns of those birds; they often visualize the data by phylogenetic trees and related methods. Your own phylogenetic analysis will consist of comparing bacterial communities in two distinct bird populations. (We admit, not as flashy as studying the flu!) | |
| − | + | Recent evidence in the burgeoning field of the microbiome suggests that the internal host environment may be a significant factor in the susceptibility/resistance profiles of individuals, populations, or species to disease pathogens. Different species of birds, although in the same family or genera taxonomically, may utilize vastly different environments and travel through flyways that are separated by thousands of miles. The associated microbial communities and potential pathogens these species encounter and carry may therefore be different, and even be a significant reason why one species is a host for a given pathogen and the other is not. As a part of Module 1, we will utilize an early approach to community profiling to gain a snapshot of the differences between gull populations in South Boston: we'll have access to samples from male and female ring-billed gulls collected at Carson Beach, and samples from male herring gulls collected at the South Bay parking lot. Should we expect these birds to be equivalent hosts for viruses? For fungi? For the bacteria that we will study? | |
| − | + | To identify the bacteria cohabiting with individual birds, we'll preferentially extract microbial (rather than animal) DNA from bird cloacal swabs and sequence a conserved region. Genes encoding ribosomal RNA (rRNA) are excellent candidates for this strategy: they are essential for life and thus the organism is unlikely to survive rRNA gene mutation. Depending on the pathogen, the small or large subunit or the internal transcribed sequence (ITS) might be the most reliable sequence for identification. We will use primers that amplify a large fragment, or amplicon, of the 16S rRNA gene sequence; this rRNA is part of the 30S small subunit. The clinical potential of 16S rRNA-based sequencing for bacterial infections is described in the [http://www.ncbi.nlm.nih.gov/pubmed/15489351 '''linked review'''] by Dr. Jill Clarridge. | |
| − | + | We must amplify the 16S rRNA gene sequences in a polymerase chain reaction (PCR), both to isolate the DNA of interest and to prepare large quantities of it. The PCR will result in a pool of 16S sequences representing different species of bacteria present approximately in proportion to their composition in the bird stool. That is, a species that is abundantly present in stool is more likely to have its DNA amplified during the PCR than is a low concentration species. The pool of 16S fragments can be cloned into a DNA vector and transformed into laboratory bacteria to produce isolated colonies. Each colony should contain identical copies of 16S DNA from a single species of bacteria, thus allowing us to deconvolve our pool of DNA and analyze individual sequences. How? Briefly, we'll perform a second DNA extraction from each of several colonies and determine the sequences through a method that resembles PCR. The individual steps will make more sense as we complete each of them, and an overview of the process as a whole is shown below. | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | [[Image: | + | [[Image:S14-M1 experiment-overview.jpg|thumb|450px|center|'''Bird gut microbiota experiment overview.''']] |
<br style="clear:both;"/> | <br style="clear:both;"/> | ||
| − | Returning to today's specific work, each of you will extract a DNA pool from a single bird | + | Returning to today's specific work, each of you will extract a DNA pool from a single bird cloacal sample using a commercial kit. Unlike most mammals, which have different excretory paths for urine and feces, birds have both their urine and feces excreted through a single opening called the cloaca. The cloacal swab is a somewhat vexing material from which to extract DNA, because many enzyme inhibitors (including materials that inhibit polymerase) are present. As described in the [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC167874/pdf/621102.pdf '''paper by Carol Kreader'''], inhibitors in feces include bile salts and environmental inhibitors such as humic compounds present in water and dirt. Chemicals that degrade DNA may be present, which is especially troubling when one wants to amplify a low concentration DNA. The DNA extraction kit contains two reagents that degrade or bind up inhibitors ��� buffer ASL and the InhibiTEX tablets ��� but unfortunately their exact contents and mechanisms of action are proprietary. |
| − | Many reagents and steps in the purification are transparent in their composition and mechanism. For example, we | + | Many reagents and steps in the purification ''are'' transparent in their composition and mechanism. For example, we know that ASL is a lysis reagent, and that we can increase the ratio of microbial:animal DNA recovered by performing lysis at a high temperature such as 70 °C. We also know that proteinase K is used to digest proteins, some of which themselves may act as PCR inhibitors or nucleases. After initial digestion and binding, the samples are purified on silica spin columns, which are also well understood. |
| − | DNA molecules passing through a silica (SiO2) column can be selectively retained by both chemical (e.g., charge) and physical (i.e., size) interactions with the porous, high surface area beads. When nucleic acids are diluted in a high concentration | + | DNA molecules passing through a silica (SiO2) column can be selectively retained by both chemical (e.g., charge) and physical (i.e., size) interactions with the porous, high surface area beads. When nucleic acids are diluted in a high concentration chaotropic salt buffer, they will tend to bind to the silica. This is because chaotropic salts (such as guanidine isothiocyanate, present in buffers AL and AW1) disrupt hydrogen-bond organization between water and macromolecules, essentially dehydrating the nucleic acids and causing them to bind to the resin. Ethanol (present in high concentrations in buffers AW1 and AW2) further precipitates the nucleic acids. The column-bound acids are washed with various buffers to remove salts and other contaminants before finally eluting in an ethanol-free, low-salt buffer in which nucleic acids are highly soluble. This final buffer is also at an increased pH to increase charge repulsion between silica and DNA that was previously screened under high salt, low pH conditions. The exact pore size and surface chemistry of the silica beads determine what sizes and kinds of nucleic acid will be bound versus washed away. After purification, two additional steps – which we'll discuss next time – can improve downstream performance in PCR. |
| − | + | We���ve made two minor modifications to the manufacturer's protocol today, due to the nature of our samples and our particular research question, respectively. One is using a somewhat lower volume of sample than recommended because our bird coalcal sample is more concentrated than the human stool the kit was designed for. The second is using chitinase in addition to proteinase K, and completing the enzymatic digestion at a lower temperature and for a longer time than recommended. Adding chitinase is useful for opening up pathogens with chitin walls, including the fungus microsporidia. More about the potential utility of this step next time! | |
| − | + | ||
| − | + | ||
==Protocols== | ==Protocols== | ||
| − | ===Part 1: DNA extraction from bird | + | ===Part 1: Begin DNA extraction from bird cloacal swab=== |
| − | The following protocol requires many tube changes. Be sure that each tube is clearly labeled with your sample number to avoid swapping samples with | + | The following protocol requires many tube changes. Be sure that each tube is clearly labeled with your sample number and initials to avoid swapping samples with anyone. There are a few other steps you can take to avoid cross-contamination: switch pipet tips at every step, keep only one tube open at a time, and avoid getting liquids on the lip of any tube or column. |
| − | Before beginning this protocol, check the maximum spin speed of your centrifuge. Some centrifuges reach 20,000 g and others reach only 16,000 g, and the time for centrifugation will have to be adjusted proportionally. Note that ''rpm'' stands for rotations per minute while ''rcf'' stands for | + | Before beginning this protocol, check the maximum spin speed of your centrifuge. Some centrifuges reach 20,000 g and others reach only 16,000 g, and the time for centrifugation will have to be adjusted proportionally. Note that ''rpm'' stands for rotations per minute while ''rcf'' stands for relative centrifugal force. It is ''rcf'' that is equivalent to g-force, not ''rpm'', because differently sized rotors will impart different forces at the same rotational speed. |
| − | #Obtain your 100 μL bird | + | #Obtain your 100 μL bird cloacal sample from the ice bucket nearest you, according to the number you are assigned on today's Talk page. |
#*Work quickly and keep the sample on your ice bucket until you have finished adding the lysis reagent. | #*Work quickly and keep the sample on your ice bucket until you have finished adding the lysis reagent. | ||
#Immediately add 1.4 mL of the lysis reagent (called buffer ASL) and vortex for ~ 1 min, until the solution is homogeneous. | #Immediately add 1.4 mL of the lysis reagent (called buffer ASL) and vortex for ~ 1 min, until the solution is homogeneous. | ||
| Line 57: | Line 51: | ||
#Incubate 1 min longer on a tube stand (with no shaking), and then centrifuge for 5 min (20K g) or 6 min (16K g). | #Incubate 1 min longer on a tube stand (with no shaking), and then centrifuge for 5 min (20K g) or 6 min (16K g). | ||
#Transfer the supernatant (usually about 500 μL) to a '''1.5 mL tube''' and again centrifuge 3 or 4 min as needed. | #Transfer the supernatant (usually about 500 μL) to a '''1.5 mL tube''' and again centrifuge 3 or 4 min as needed. | ||
| − | #In a fresh '''1.5 mL tube''', dispense 15 μL proteinase K. Only then should you add 200 μL of the supernatant | + | #In a fresh '''1.5 mL tube''', dispense 15 μL proteinase K. Only then should you add 200 μL of the supernatant from step 9 followed by 200 μL of buffer AL, pipetting to mix each time. Finally, add 15 μL chitinase. |
| − | #Vortex for 15 sec (until solution is homogeneous) and incubate in the 56 °C oven for about | + | #Vortex for 15 sec (until solution is homogeneous) and incubate in the 56 °C oven for about 1.5 hours. |
===Part 2: Lab practical=== | ===Part 2: Lab practical=== | ||
| Line 64: | Line 58: | ||
You and your partner may work together on the lab practical. (Note: this collaboration will not be the case for future quizzes.) You are of course welcome to give different answers should you disagree. | You and your partner may work together on the lab practical. (Note: this collaboration will not be the case for future quizzes.) You are of course welcome to give different answers should you disagree. | ||
| − | ===Part 3: DNA extraction from bird | + | Afterward, you can work on your notebook and start preparing for Part 3. |
| + | |||
| + | ===Part 3: Complete DNA extraction from bird cloacal swab=== | ||
#Quick-spin to recover the part of the sample that has condensed in the eppendorf tube lid. | #Quick-spin to recover the part of the sample that has condensed in the eppendorf tube lid. | ||
#*To quick-spin, hold down the "short" button on your centrifuge for 3-5 seconds, then release. | #*To quick-spin, hold down the "short" button on your centrifuge for 3-5 seconds, then release. | ||
| − | #Add 200 μL of ethanol, mix by briefly vortexing, and transfer to a QIAamp spin column (atop its 2 mL collection tube). | + | #Add 200 μL of ethanol, mix by briefly vortexing, and transfer to a QIAamp spin column (atop its 2 mL capless collection tube). |
| − | #*Be sure to label the column | + | #*Be sure to label the column rather than the tube with your sample number and initials. |
| − | #Centrifuge for 1 min (1.5 min on slower centrifuges), and if necessary repeat the centrifugation until all the sample has gone through the column. (Unlikely in our case | + | #Centrifuge for 1 min (1.5 min on slower centrifuges), and if necessary repeat the centrifugation until all the sample has gone through the column. (Unlikely in our case). |
| − | #Move the spin column to a fresh collection tube, add 500 μL buffer AW1, and spin 1 or 1.5 min as needed. | + | #Move the spin column to a fresh capless collection tube, add 500 μL buffer AW1, and spin 1 or 1.5 min as needed. |
#Move the spin column to a fresh collection tube, add 500 μL buffer AW2, and spin 3 or 4 min as needed. | #Move the spin column to a fresh collection tube, add 500 μL buffer AW2, and spin 3 or 4 min as needed. | ||
| − | #In the meantime, trim the cap off a fresh 1.5 mL eppendorf tube using small scissors that have been wiped down with 70% ethanol. Prepare a sticky label (in your team color) for the top: write the date | + | #'''In the meantime''', trim the cap off a fresh 1.5 mL eppendorf tube using small scissors that have been wiped down with 70% ethanol. Save the cap! Prepare a sticky label (in your team color) for the top: write the date, your sample ID, and your initials. You should also label the ''side'' of each tube, at least with a short unique identifier, so you don't lose track of which sample is which in the following step. |
#Move to yet another fresh collection tube and spin 1 or 1.5 min to rid residual buffer. | #Move to yet another fresh collection tube and spin 1 or 1.5 min to rid residual buffer. | ||
| − | #*This step completely removes remaining ethanol that could interfere with future | + | #*This step completely removes remaining ethanol that could interfere with future steps. |
#Now transfer the column to your trimmed, well-labeled 1.5 mL tube and carefully pipet 150 μL buffer of AE onto the membrane. | #Now transfer the column to your trimmed, well-labeled 1.5 mL tube and carefully pipet 150 μL buffer of AE onto the membrane. | ||
| − | #Incubate for 5 min, then spin for 1 or 1.5 min, cap, and store in your ice bucket. We will collect each sample and store it at -20 °C until next time. | + | #Incubate for 5 min, then spin for 1 or 1.5 min, cap, and store in your ice bucket. We will collect each sample and store it at -20 °C until next week. |
| + | |||
| + | ==For next time== | ||
| + | |||
| + | #Continue to familiarize yourself with the workings of OpenWetWare. Specifically, you should: | ||
| + | #*Create a user page, add some content to it, and add it to the '''[[20.109(S14):People]]''' page. Be careful about how you put your e-mail address on your page; make sure to spell out "at" or otherwise break your address up so it can't be scraped up by spambots. It is fine to use the sample OWW user page as a starting point, but be sure to erase or replace information that does not pertain to you (such as someone else's publications!) and erase or replace general content such as "Research Interest 1" with your own interests. | ||
| + | #*Complete the student registration/questionnaire from [http://openwetware.org/wiki/20.109(S14):Student_registration_and_questionnaire | '''this link'''] to turn in next time. Click on the "edit" tab of this page, then copy the "source code" you see to your user page, fill out the form, and print it. You do not need to keep the information on your user page after printing it. | ||
| + | #The major assessment for this module will be a formal written summary. Start to familiarize yourself with its expected structure and content at [[20.109%28S14%29:Microbiome_summary | '''this link''']]. | ||
| + | #One week from today [[20.109(S14):16S PCR and paper discussion (Day3)#Parts 2 + 3: Journal article discussion and WAC session| '''(Mod 1, Day 3)''']], we will have an in-class discussion about an article from the primary scientific literature. You might begin reading and thinking about the paper now, rather than trying to do so only in the two days between Day 2 and Day 3. The paper is linked from PubMed [http://www.ncbi.nlm.nih.gov/pubmed/20668239 '''here''']. | ||
| + | #Finish your EHS training if you haven't already. | ||
| + | |||
| + | ==Reagent list== | ||
| + | |||
| + | * QIAamp DNA Stool Mini Kit from Qiagen | ||
| + | * Chitinase from Sigma (NOT part of usual Qiagen kit!) | ||
| + | **Stock solution prepared at 0.02666 U/μL concentration in 50 mM potassium phosphate | ||
| + | **Therefore 0.4 U are added to each extraction | ||
| + | * Ethanol (200 proof) | ||
| + | |||
| + | ==Navigation Links== | ||
| + | Next Day: [[20.109(S14):Diagnostic primer design (Day2) | Diagnostic primer design]] | ||
| + | Previous Day: [[20.109(S14):Lab tour | Orientation]] | ||
Latest revision as of 13:58, 29 July 2015
Contents
Introduction
Please begin by reading the Module 1 overview.
Today we'll begin our primary experiment, a phylogenetic analysis of bird gut microbiota. Through birds' stool, their resident microbes can be transferred to the environment and in some cases infect other animals. Perhaps the most well-known pathogenic avian microbe with zoonotic potential (potential for inter-species transmission) is the flu virus. For your safety, all the samples we will work with have been screened to exclude those carrying human���pathogenic flu strains. However, we will be able to mine much of the same intellectual content that we could were we studying flu directly.
Investigations in disease ecology, or "the ecological study of host-pathogen interactions within the context of their environment and evolution" (via Scitable), help determine how pathogens transmit and cause disease, persist, and evolve in host organisms as different as humans and birds. Pathogen, host, and environment all play roles in defining the natural history of disease. In studies of pathogens that cause zoonotic disease, researchers are particularly interested in defining major influences on pathogen distribution, transmission, and evolution. The Runstadler lab currently studies the disease ecology of influenza viruses in several groups of birds and other animals (see for example this paper and this one), a research area that can provide information useful for predicting the next flu pandemic and designing effective vaccines that will be produced in time. Researchers track viral mutations/evolution, infection of different bird species (including co-infection by multiple strains), and the trafficking patterns of those birds; they often visualize the data by phylogenetic trees and related methods. Your own phylogenetic analysis will consist of comparing bacterial communities in two distinct bird populations. (We admit, not as flashy as studying the flu!)
Recent evidence in the burgeoning field of the microbiome suggests that the internal host environment may be a significant factor in the susceptibility/resistance profiles of individuals, populations, or species to disease pathogens. Different species of birds, although in the same family or genera taxonomically, may utilize vastly different environments and travel through flyways that are separated by thousands of miles. The associated microbial communities and potential pathogens these species encounter and carry may therefore be different, and even be a significant reason why one species is a host for a given pathogen and the other is not. As a part of Module 1, we will utilize an early approach to community profiling to gain a snapshot of the differences between gull populations in South Boston: we'll have access to samples from male and female ring-billed gulls collected at Carson Beach, and samples from male herring gulls collected at the South Bay parking lot. Should we expect these birds to be equivalent hosts for viruses? For fungi? For the bacteria that we will study?
To identify the bacteria cohabiting with individual birds, we'll preferentially extract microbial (rather than animal) DNA from bird cloacal swabs and sequence a conserved region. Genes encoding ribosomal RNA (rRNA) are excellent candidates for this strategy: they are essential for life and thus the organism is unlikely to survive rRNA gene mutation. Depending on the pathogen, the small or large subunit or the internal transcribed sequence (ITS) might be the most reliable sequence for identification. We will use primers that amplify a large fragment, or amplicon, of the 16S rRNA gene sequence; this rRNA is part of the 30S small subunit. The clinical potential of 16S rRNA-based sequencing for bacterial infections is described in the linked review by Dr. Jill Clarridge.
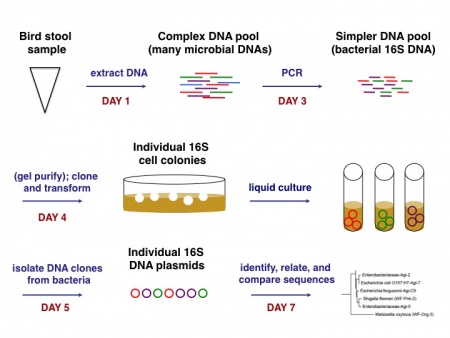
We must amplify the 16S rRNA gene sequences in a polymerase chain reaction (PCR), both to isolate the DNA of interest and to prepare large quantities of it. The PCR will result in a pool of 16S sequences representing different species of bacteria present approximately in proportion to their composition in the bird stool. That is, a species that is abundantly present in stool is more likely to have its DNA amplified during the PCR than is a low concentration species. The pool of 16S fragments can be cloned into a DNA vector and transformed into laboratory bacteria to produce isolated colonies. Each colony should contain identical copies of 16S DNA from a single species of bacteria, thus allowing us to deconvolve our pool of DNA and analyze individual sequences. How? Briefly, we'll perform a second DNA extraction from each of several colonies and determine the sequences through a method that resembles PCR. The individual steps will make more sense as we complete each of them, and an overview of the process as a whole is shown below.
Returning to today's specific work, each of you will extract a DNA pool from a single bird cloacal sample using a commercial kit. Unlike most mammals, which have different excretory paths for urine and feces, birds have both their urine and feces excreted through a single opening called the cloaca. The cloacal swab is a somewhat vexing material from which to extract DNA, because many enzyme inhibitors (including materials that inhibit polymerase) are present. As described in the paper by Carol Kreader, inhibitors in feces include bile salts and environmental inhibitors such as humic compounds present in water and dirt. Chemicals that degrade DNA may be present, which is especially troubling when one wants to amplify a low concentration DNA. The DNA extraction kit contains two reagents that degrade or bind up inhibitors ��� buffer ASL and the InhibiTEX tablets ��� but unfortunately their exact contents and mechanisms of action are proprietary.
Many reagents and steps in the purification are transparent in their composition and mechanism. For example, we know that ASL is a lysis reagent, and that we can increase the ratio of microbial:animal DNA recovered by performing lysis at a high temperature such as 70 °C. We also know that proteinase K is used to digest proteins, some of which themselves may act as PCR inhibitors or nucleases. After initial digestion and binding, the samples are purified on silica spin columns, which are also well understood.
DNA molecules passing through a silica (SiO2) column can be selectively retained by both chemical (e.g., charge) and physical (i.e., size) interactions with the porous, high surface area beads. When nucleic acids are diluted in a high concentration chaotropic salt buffer, they will tend to bind to the silica. This is because chaotropic salts (such as guanidine isothiocyanate, present in buffers AL and AW1) disrupt hydrogen-bond organization between water and macromolecules, essentially dehydrating the nucleic acids and causing them to bind to the resin. Ethanol (present in high concentrations in buffers AW1 and AW2) further precipitates the nucleic acids. The column-bound acids are washed with various buffers to remove salts and other contaminants before finally eluting in an ethanol-free, low-salt buffer in which nucleic acids are highly soluble. This final buffer is also at an increased pH to increase charge repulsion between silica and DNA that was previously screened under high salt, low pH conditions. The exact pore size and surface chemistry of the silica beads determine what sizes and kinds of nucleic acid will be bound versus washed away. After purification, two additional steps – which we'll discuss next time – can improve downstream performance in PCR.
We���ve made two minor modifications to the manufacturer's protocol today, due to the nature of our samples and our particular research question, respectively. One is using a somewhat lower volume of sample than recommended because our bird coalcal sample is more concentrated than the human stool the kit was designed for. The second is using chitinase in addition to proteinase K, and completing the enzymatic digestion at a lower temperature and for a longer time than recommended. Adding chitinase is useful for opening up pathogens with chitin walls, including the fungus microsporidia. More about the potential utility of this step next time!
Protocols
Part 1: Begin DNA extraction from bird cloacal swab
The following protocol requires many tube changes. Be sure that each tube is clearly labeled with your sample number and initials to avoid swapping samples with anyone. There are a few other steps you can take to avoid cross-contamination: switch pipet tips at every step, keep only one tube open at a time, and avoid getting liquids on the lip of any tube or column.
Before beginning this protocol, check the maximum spin speed of your centrifuge. Some centrifuges reach 20,000 g and others reach only 16,000 g, and the time for centrifugation will have to be adjusted proportionally. Note that rpm stands for rotations per minute while rcf stands for relative centrifugal force. It is rcf that is equivalent to g-force, not rpm, because differently sized rotors will impart different forces at the same rotational speed.
- Obtain your 100 μL bird cloacal sample from the ice bucket nearest you, according to the number you are assigned on today's Talk page.
- Work quickly and keep the sample on your ice bucket until you have finished adding the lysis reagent.
- Immediately add 1.4 mL of the lysis reagent (called buffer ASL) and vortex for ~ 1 min, until the solution is homogeneous.
- The fastest way to add the appropriate amount of ASL is to add 0.7 mL twice with your P1000; that way you don't have to rotate the pipet setting in between additions.
- A few insoluble particulates may remain. Try vortexing for another 20-30 sec interval up to four more times, and stop vortexing when the sample no longer visibly changes over that interval.
- Heat at 70 °C for 5 min on the heat block at the front bench.
- Vortex for 15 sec and centrifuge for 1 min at 20,000 rcf or 1.5 min at 16,000 rcf. Place your tubes so that weight is equally distributed in the centrifuge.
- Unfortunately, your centrifuges cannot be set for 1.25 min exactly.
- Transfer 1.2 mL of supernatant into a fresh 2 mL tube.
- Be sure to use the special 2 mL eppendorfs here and not the standard 1.5 mL eppendorf tubes.
- Hold the foil-covered InhibitEX tablet over the tube, and gently push until the tablet pierces through the foil and falls into the tube.
- Vortex until completely dissolved, which takes about 3 min for these samples.
- The solution will be homogeneous but somewhat thick.
- Incubate 1 min longer on a tube stand (with no shaking), and then centrifuge for 5 min (20K g) or 6 min (16K g).
- Transfer the supernatant (usually about 500 μL) to a 1.5 mL tube and again centrifuge 3 or 4 min as needed.
- In a fresh 1.5 mL tube, dispense 15 μL proteinase K. Only then should you add 200 μL of the supernatant from step 9 followed by 200 μL of buffer AL, pipetting to mix each time. Finally, add 15 μL chitinase.
- Vortex for 15 sec (until solution is homogeneous) and incubate in the 56 °C oven for about 1.5 hours.
Part 2: Lab practical
You and your partner may work together on the lab practical. (Note: this collaboration will not be the case for future quizzes.) You are of course welcome to give different answers should you disagree.
Afterward, you can work on your notebook and start preparing for Part 3.
Part 3: Complete DNA extraction from bird cloacal swab
- Quick-spin to recover the part of the sample that has condensed in the eppendorf tube lid.
- To quick-spin, hold down the "short" button on your centrifuge for 3-5 seconds, then release.
- Add 200 μL of ethanol, mix by briefly vortexing, and transfer to a QIAamp spin column (atop its 2 mL capless collection tube).
- Be sure to label the column rather than the tube with your sample number and initials.
- Centrifuge for 1 min (1.5 min on slower centrifuges), and if necessary repeat the centrifugation until all the sample has gone through the column. (Unlikely in our case).
- Move the spin column to a fresh capless collection tube, add 500 μL buffer AW1, and spin 1 or 1.5 min as needed.
- Move the spin column to a fresh collection tube, add 500 μL buffer AW2, and spin 3 or 4 min as needed.
- In the meantime, trim the cap off a fresh 1.5 mL eppendorf tube using small scissors that have been wiped down with 70% ethanol. Save the cap! Prepare a sticky label (in your team color) for the top: write the date, your sample ID, and your initials. You should also label the side of each tube, at least with a short unique identifier, so you don't lose track of which sample is which in the following step.
- Move to yet another fresh collection tube and spin 1 or 1.5 min to rid residual buffer.
- This step completely removes remaining ethanol that could interfere with future steps.
- Now transfer the column to your trimmed, well-labeled 1.5 mL tube and carefully pipet 150 μL buffer of AE onto the membrane.
- Incubate for 5 min, then spin for 1 or 1.5 min, cap, and store in your ice bucket. We will collect each sample and store it at -20 °C until next week.
For next time
- Continue to familiarize yourself with the workings of OpenWetWare. Specifically, you should:
- Create a user page, add some content to it, and add it to the 20.109(S14):People page. Be careful about how you put your e-mail address on your page; make sure to spell out "at" or otherwise break your address up so it can't be scraped up by spambots. It is fine to use the sample OWW user page as a starting point, but be sure to erase or replace information that does not pertain to you (such as someone else's publications!) and erase or replace general content such as "Research Interest 1" with your own interests.
- Complete the student registration/questionnaire from | this link to turn in next time. Click on the "edit" tab of this page, then copy the "source code" you see to your user page, fill out the form, and print it. You do not need to keep the information on your user page after printing it.
- The major assessment for this module will be a formal written summary. Start to familiarize yourself with its expected structure and content at this link.
- One week from today (Mod 1, Day 3), we will have an in-class discussion about an article from the primary scientific literature. You might begin reading and thinking about the paper now, rather than trying to do so only in the two days between Day 2 and Day 3. The paper is linked from PubMed here.
- Finish your EHS training if you haven't already.
Reagent list
- QIAamp DNA Stool Mini Kit from Qiagen
- Chitinase from Sigma (NOT part of usual Qiagen kit!)
- Stock solution prepared at 0.02666 U/μL concentration in 50 mM potassium phosphate
- Therefore 0.4 U are added to each extraction
- Ethanol (200 proof)
Next Day: Diagnostic primer design
Previous Day: Orientation